Strip charts: 1-D scatter plots - R Base Graphs
Previously, we described the essentials of R programming and provided quick start guides for importing data into R.
Here, we’ll describe how to create strip charts (i.e., one dimensional scatter plots or dot plots) in R. These plots are a good alternative to box plots when sample sizes are small.
Pleleminary tasks
Launch RStudio as described here: Running RStudio and setting up your working directory
Prepare your data as described here: Best practices for preparing your data and save it in an external .txt tab or .csv files
Import your data into R as described here: Fast reading of data from txt|csv files into R: readr package.
Here, we’ll use the R built-in ToothGrowth data set.
ToothGrowth$dose <- as.factor(ToothGrowth$dose)
# Print the first 6 rows
head(ToothGrowth, 6)## len supp dose
## 1 4.2 VC 0.5
## 2 11.5 VC 0.5
## 3 7.3 VC 0.5
## 4 5.8 VC 0.5
## 5 6.4 VC 0.5
## 6 10.0 VC 0.5R base function: stripchart()
stripchart(x, data = NULL method = "overplot", jitter = 0.1)- x: the data from which the plots are to be produced. Allowed values are one or a list of numeric vector, each corresponding to a component plot.
- data: a data.frame (or list) from which the variables in x should be taken.
- Method: the method to be used to separate coincident points. Allowed values are one of “overplot”, “jitter” or “stack”.
- jitter: when method = “jitter” is used, jitter gives the amount of jittering applied.
Create strip charts
# Plot len by dose
stripchart(len ~ dose, data = ToothGrowth,
pch = 19, frame = FALSE)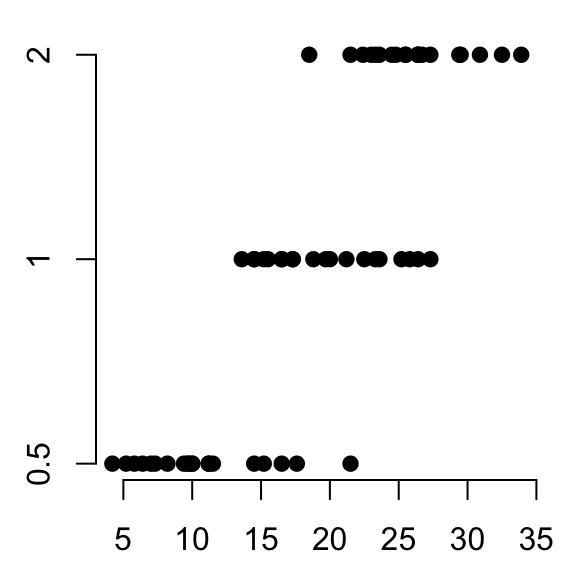
# Vertical plot using method = "jitter"
stripchart(len ~ dose, data = ToothGrowth,
pch = 19, frame = FALSE, vertical = TRUE,
method = "jitter")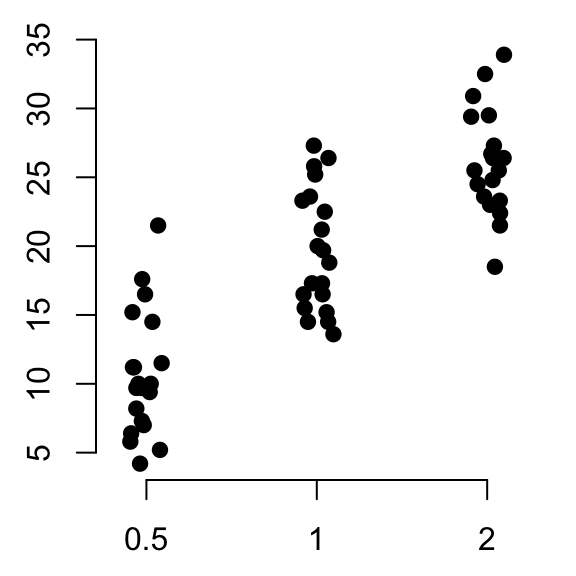
# Change point shapes (pch) and colors by groups
# add main title and axis labels
stripchart(len ~ dose, data = ToothGrowth,
frame = FALSE, vertical = TRUE,
method = "jitter", pch = c(21, 18, 16),
col = c("#999999", "#E69F00", "#56B4E9"),
main = "Length by dose", xlab = "Dose", ylab = "Length")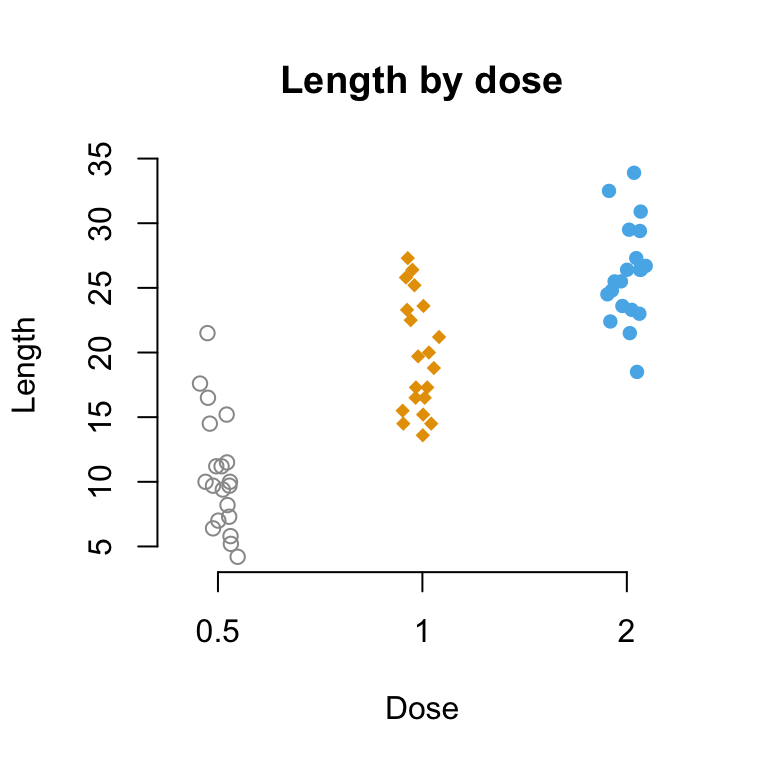
See also
Infos
This analysis has been performed using R statistical software (ver. 3.2.4).
Enjoyed this article? I’d be very grateful if you’d help it spread by emailing it to a friend, or sharing it on Twitter, Facebook or Linked In.
Show me some love with the like buttons below... Thank you and please don't forget to share and comment below!!
Show me some love with the like buttons below... Thank you and please don't forget to share and comment below!!
Avez vous aimé cet article? Je vous serais très reconnaissant si vous aidiez à sa diffusion en l'envoyant par courriel à un ami ou en le partageant sur Twitter, Facebook ou Linked In.
Montrez-moi un peu d'amour avec les like ci-dessous ... Merci et n'oubliez pas, s'il vous plaît, de partager et de commenter ci-dessous!
Montrez-moi un peu d'amour avec les like ci-dessous ... Merci et n'oubliez pas, s'il vous plaît, de partager et de commenter ci-dessous!
Recommended for You!
Recommended for you
This section contains the best data science and self-development resources to help you on your path.
Books - Data Science
Our Books
- Practical Guide to Cluster Analysis in R by A. Kassambara (Datanovia)
- Practical Guide To Principal Component Methods in R by A. Kassambara (Datanovia)
- Machine Learning Essentials: Practical Guide in R by A. Kassambara (Datanovia)
- R Graphics Essentials for Great Data Visualization by A. Kassambara (Datanovia)
- GGPlot2 Essentials for Great Data Visualization in R by A. Kassambara (Datanovia)
- Network Analysis and Visualization in R by A. Kassambara (Datanovia)
- Practical Statistics in R for Comparing Groups: Numerical Variables by A. Kassambara (Datanovia)
- Inter-Rater Reliability Essentials: Practical Guide in R by A. Kassambara (Datanovia)
Others
- R for Data Science: Import, Tidy, Transform, Visualize, and Model Data by Hadley Wickham & Garrett Grolemund
- Hands-On Machine Learning with Scikit-Learn, Keras, and TensorFlow: Concepts, Tools, and Techniques to Build Intelligent Systems by Aurelien Géron
- Practical Statistics for Data Scientists: 50 Essential Concepts by Peter Bruce & Andrew Bruce
- Hands-On Programming with R: Write Your Own Functions And Simulations by Garrett Grolemund & Hadley Wickham
- An Introduction to Statistical Learning: with Applications in R by Gareth James et al.
- Deep Learning with R by François Chollet & J.J. Allaire
- Deep Learning with Python by François Chollet
Get involved :
Click to follow us on Facebook :
Comment this article by clicking on "Discussion" button (top-right position of this page)
Click to follow us on Facebook :
Comment this article by clicking on "Discussion" button (top-right position of this page)







