Installing and Using R Packages
In our previous articles, we published i) guides for installing and launching R/RStudio, ii) the basics of R programming, and ii) guides for finding help in R.
Here, we’ll describe:
- what is an R package
- and how to install and use R packages
What is R packages?
An R package is an extension of R containing data sets and specific functions to solve specific questions.
R comes with standard (or base) packages, which contain the basic functions and data sets as well as standard statistical and graphical functions that allow R to work.
There are also thousands other R packages available for download and installation from CRAN, Bioconductor and GitHub repositories.
After installation, you must first load the package for using the functions in the package.
Installing R packages
Packages can be installed either from CRAN (for general packages), from Bioconductor (for biology-related packages) or from Github (developing versions of packages).
Install a package from CRAN
The function install.packages() is used to install a package from CRAN. The syntax is as follow:
install.packages("package_name")For example, to install the package named readr, type this:
install.packages("readr")Note that, every time you install an R package, R may ask you to specify a CRAN mirror (or server). Choose one that’s close to your location, and R will connect to that server to download and install the package files.
It’s also possible to install multiple packages at the same time, as follow:
install.packages(c("readr", "ggplot2"))Install a package from Bioconductor
Bioconductor contains packages for analyzing biological related data. In the following R code, we want to install the R/Bioconductor package limma, which is dedicated to analyse genomic data.
To install a package from Bioconductor, use this:
source("https://bioconductor.org/biocLite.R")
biocLite("limma")Install a package from Github
GitHub is a repository useful for all software development and data analysis, including R packages. It makes sharing your package easy. You can read more about GitHub here: Git and GitHub, by Hadley Wickham.
To install a package from GitHub, the R package devtools (by Hadley Wickham) can be used. You should first install devtools if you don’t have it installed on your computer.
For example, the following R code installs the latest version of survminer R package developed by A. Kassambara (https://github.com/kassambara/survminer).
install.packages("devtools")
devtools::install_github("kassambara/survminer")View the list of installed packages
To view the list of the already installed packages on your computer, type :
installed.packages()Note that, in RStudio, the list of installed packages are available in the lower right window under Packages tab (see the image below).
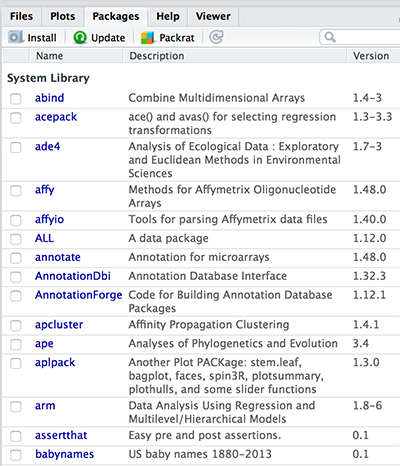
Folder containing installed packages
R packages are installed in a directory called library. The R function .libPaths() can be used to get the path to the library.
.libPaths()[1] "/Library/Frameworks/R.framework/Versions/3.2/Resources/library"Load and use an R package
To use a specific function available in an R package, you have to load the R package using the function library().
In the following R code, we want to import a file (“https://www.sthda.com/upload/decathlon.txt”) into R using the R package readr, which has been installed in the previous section.
The function read_tsv() [in readr] can be used to import a tab separated .txt file:
# Import my data
library("readr")
my_data <- read_tsv("https://www.sthda.com/upload/decathlon.txt")
# View the first 6 rows and tge first 6 columns
# syntax: my_data[row, column]
my_data[1:6, 1:6] name 100m Long.jump Shot.put High.jump 400m
1 SEBRLE 11.04 7.58 14.83 2.07 49.81
2 CLAY 10.76 7.40 14.26 1.86 49.37
3 KARPOV 11.02 7.30 14.77 2.04 48.37
4 BERNARD 11.02 7.23 14.25 1.92 48.93
5 YURKOV 11.34 7.09 15.19 2.10 50.42
6 WARNERS 11.11 7.60 14.31 1.98 48.68View loaded R packages
To view the list of loaded (or attached) packages during an R session, use the function search():
search() [1] ".GlobalEnv" "package:readr" "package:stats" "package:graphics"
[5] "package:grDevices" "package:utils" "package:datasets" "package:methods"
[9] "Autoloads" "package:base" If you’re done with the package readr and you want to unload it, use the function detach():
detach("readr", unload = TRUE)Remove installed packages
To remove an installed R package, use the function remove.packages() as follow:
remove.packages("package_name")Update installed packages
If you want to update all installed R packages, type this:
update.packages()To update specific installed packages, say readr and ggplot2, use this:
update.packages(oldPkgs = c("readr", "ggplot2"))Summary
install.packages(“package_name”): Install a package
library(“package_name”): Load and use a package
detach(“package_name”, unload = TRUE): Unload a package
remove.packages(“package_name”): Remove an installed package from your computer
- update.packages(oldPkgs = “package_name”): Update a package
Infos
This analysis has been performed using R software (ver. 3.2.3).
Show me some love with the like buttons below... Thank you and please don't forget to share and comment below!!
Montrez-moi un peu d'amour avec les like ci-dessous ... Merci et n'oubliez pas, s'il vous plaît, de partager et de commenter ci-dessous!
Recommended for You!
Recommended for you
This section contains the best data science and self-development resources to help you on your path.
Books - Data Science
Our Books
- Practical Guide to Cluster Analysis in R by A. Kassambara (Datanovia)
- Practical Guide To Principal Component Methods in R by A. Kassambara (Datanovia)
- Machine Learning Essentials: Practical Guide in R by A. Kassambara (Datanovia)
- R Graphics Essentials for Great Data Visualization by A. Kassambara (Datanovia)
- GGPlot2 Essentials for Great Data Visualization in R by A. Kassambara (Datanovia)
- Network Analysis and Visualization in R by A. Kassambara (Datanovia)
- Practical Statistics in R for Comparing Groups: Numerical Variables by A. Kassambara (Datanovia)
- Inter-Rater Reliability Essentials: Practical Guide in R by A. Kassambara (Datanovia)
Others
- R for Data Science: Import, Tidy, Transform, Visualize, and Model Data by Hadley Wickham & Garrett Grolemund
- Hands-On Machine Learning with Scikit-Learn, Keras, and TensorFlow: Concepts, Tools, and Techniques to Build Intelligent Systems by Aurelien Géron
- Practical Statistics for Data Scientists: 50 Essential Concepts by Peter Bruce & Andrew Bruce
- Hands-On Programming with R: Write Your Own Functions And Simulations by Garrett Grolemund & Hadley Wickham
- An Introduction to Statistical Learning: with Applications in R by Gareth James et al.
- Deep Learning with R by François Chollet & J.J. Allaire
- Deep Learning with Python by François Chollet
Click to follow us on Facebook :
Comment this article by clicking on "Discussion" button (top-right position of this page)







