One-Sample Wilcoxon Signed Rank Test in R
What’s one-sample Wilcoxon signed rank test?
Note that, the data should be distributed symmetrically around the median. In other words, there should be roughly the same number of values above and below the median.

Research questions and statistical hypotheses
Typical research questions are:
- whether the median (\(m\)) of the sample is equal to the theoretical value (\(m_0\))?
- whether the median (\(m\)) of the sample is less than to the theoretical value (\(m_0\))?
- whether the median (\(m\)) of the sample is greater than to the theoretical value(\(m_0\))?
In statistics, we can define the corresponding null hypothesis (\(H_0\)) as follow:
- \(H_0: m = m_0\)
- \(H_0: m \leq m_0\)
- \(H_0: m \geq m_0\)
The corresponding alternative hypotheses (\(H_a\)) are as follow:
- \(H_a: m \ne m_0\) (different)
- \(H_a: m > m_0\) (greater)
- \(H_a: m < m_0\) (less)
Note that:
- Hypotheses 1) are called two-tailed tests
- Hypotheses 2) and 3) are called one-tailed tests
Visualize your data and compute one-sample Wilcoxon test in R
Install ggpubr R package for data visualization
You can draw R base graphs as described at this link: R base graphs. Here, we’ll use the ggpubr R package for an easy ggplot2-based data visualization
- Install the latest version from GitHub as follow (recommended):
# Install
if(!require(devtools)) install.packages("devtools")
devtools::install_github("kassambara/ggpubr")- Or, install from CRAN as follow:
install.packages("ggpubr")R function to compute one-sample Wilcoxon test
To perform one-sample Wilcoxon-test, the R function wilcox.test() can be used as follow:
wilcox.test(x, mu = 0, alternative = "two.sided")- x: a numeric vector containing your data values
- mu: the theoretical mean/median value. Default is 0 but you can change it.
- alternative: the alternative hypothesis. Allowed value is one of “two.sided” (default), “greater” or “less”.
Import your data into R
Prepare your data as specified here: Best practices for preparing your data set for R
Save your data in an external .txt tab or .csv files
Import your data into R as follow:
# If .txt tab file, use this
my_data <- read.delim(file.choose())
# Or, if .csv file, use this
my_data <- read.csv(file.choose())Here, we’ll use an example data set containing the weight of 10 mice.
We want to know, if the median weight of the mice differs from 25g?
set.seed(1234)
my_data <- data.frame(
name = paste0(rep("M_", 10), 1:10),
weight = round(rnorm(10, 20, 2), 1)
)Check your data
# Print the first 10 rows of the data
head(my_data, 10) name weight
1 M_1 17.6
2 M_2 20.6
3 M_3 22.2
4 M_4 15.3
5 M_5 20.9
6 M_6 21.0
7 M_7 18.9
8 M_8 18.9
9 M_9 18.9
10 M_10 18.2# Statistical summaries of weight
summary(my_data$weight) Min. 1st Qu. Median Mean 3rd Qu. Max.
15.30 18.38 18.90 19.25 20.82 22.20 - Min.: the minimum value
- 1st Qu.: The first quartile. 25% of values are lower than this.
- Median: the median value. Half the values are lower; half are higher.
- 3rd Qu.: the third quartile. 75% of values are higher than this.
- Max.: the maximum value
Visualize your data using box plots
library(ggpubr)
ggboxplot(my_data$weight,
ylab = "Weight (g)", xlab = FALSE,
ggtheme = theme_minimal())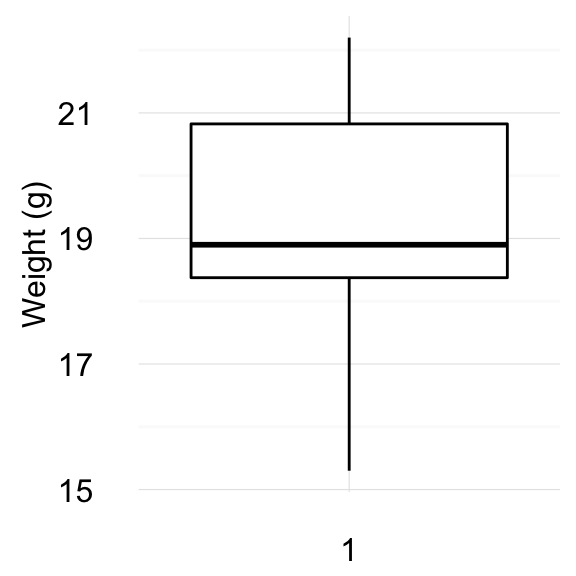
One-Sample Wilcoxon Signed Rank Test in R
Compute one-sample Wilcoxon test
We want to know, if the average weight of the mice differs from 25g (two-tailed test)?
# One-sample wilcoxon test
res <- wilcox.test(my_data$weight, mu = 25)
# Printing the results
res
Wilcoxon signed rank test with continuity correction
data: my_data$weight
V = 0, p-value = 0.005793
alternative hypothesis: true location is not equal to 25# print only the p-value
res$p.value[1] 0.005793045The p-value of the test is 0.005793, which is less than the significance level alpha = 0.05. We can reject the null hypothesis and conclude that the average weight of the mice is significantly different from 25g with a p-value = 0.005793.
Note that:
- if you want to test whether the median weight of mice is less than 25g (one-tailed test), type this:
wilcox.test(my_data$weight, mu = 25,
alternative = "less")- Or, if you want to test whether the median weight of mice is greater than 25g (one-tailed test), type this:
wilcox.test(my_data$weight, mu = 25,
alternative = "greater")See also
Infos
This analysis has been performed using R software (ver. 3.2.4).
Show me some love with the like buttons below... Thank you and please don't forget to share and comment below!!
Montrez-moi un peu d'amour avec les like ci-dessous ... Merci et n'oubliez pas, s'il vous plaît, de partager et de commenter ci-dessous!
Recommended for You!
Recommended for you
This section contains the best data science and self-development resources to help you on your path.
Books - Data Science
Our Books
- Practical Guide to Cluster Analysis in R by A. Kassambara (Datanovia)
- Practical Guide To Principal Component Methods in R by A. Kassambara (Datanovia)
- Machine Learning Essentials: Practical Guide in R by A. Kassambara (Datanovia)
- R Graphics Essentials for Great Data Visualization by A. Kassambara (Datanovia)
- GGPlot2 Essentials for Great Data Visualization in R by A. Kassambara (Datanovia)
- Network Analysis and Visualization in R by A. Kassambara (Datanovia)
- Practical Statistics in R for Comparing Groups: Numerical Variables by A. Kassambara (Datanovia)
- Inter-Rater Reliability Essentials: Practical Guide in R by A. Kassambara (Datanovia)
Others
- R for Data Science: Import, Tidy, Transform, Visualize, and Model Data by Hadley Wickham & Garrett Grolemund
- Hands-On Machine Learning with Scikit-Learn, Keras, and TensorFlow: Concepts, Tools, and Techniques to Build Intelligent Systems by Aurelien Géron
- Practical Statistics for Data Scientists: 50 Essential Concepts by Peter Bruce & Andrew Bruce
- Hands-On Programming with R: Write Your Own Functions And Simulations by Garrett Grolemund & Hadley Wickham
- An Introduction to Statistical Learning: with Applications in R by Gareth James et al.
- Deep Learning with R by François Chollet & J.J. Allaire
- Deep Learning with Python by François Chollet
Click to follow us on Facebook :
Comment this article by clicking on "Discussion" button (top-right position of this page)








