RNA sequencing data analysis - alignment and reads counting using cufflinks
This analysis was performed using R (ver. 3.1.0).Introduction
RNA-Seq is a valuable experiment for quantifying both the types and the amount of RNA molecules in a sample. The aims of this article is to show you :
- How to align the reads using tophat2?
- How to count the number of reads per features using cufflinks?
- How to visualize bam file using IGV ?
Download data
We will work with the Hammer et al dataset, as prepared by the ReCount website.
The Hammer et al paper: http://www.ncbi.nlm.nih.gov/pubmed?term=20452967 The GEO page: http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE20895 The sample I will align: http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSM539553 which points to the SRA: ftp://ftp-trace.ncbi.nlm.nih.gov/sra/sra-instant/reads/ByExp/sra/SRX%2FSRX020%2FSRX020088/SRR042499/
download the file
#download the file
wget ftp://ftp-trace.ncbi.nlm.nih.gov/sra/sra-instant/reads/ByExp/sra/SRX%2FSRX020%2FSRX020088/SRR042499/SRR042499.sra
#convert to FastQ
fastq-dump SRR042499.sraDownload reference genome from tophat website
wget wget [url=ftp://igenome:G3nom3s4u@ussd-ftp.illumina.com/Rattus_norvegicus/Ensembl/RGSC3.4/Rattus_norvegicus_Ensembl_RGSC3.4.tar.gz]ftp://igenome:G3nom3s4u@ussd-ftp.illumina.com/Rattus_norvegicus/Ensembl/RGSC3.4/Rattus_norvegicus_Ensembl_RGSC3.4.tar.gz[/url]
tar zxvf Rattus_norvegicus_Ensembl_RGSC3.4.tar.gzThe genome was downloaded from Illumina iGenomes
Align the reads
The tophat call to align the reads (this take account for splice read) :
tophat2 -o tophat_out -p 10 /path/to/Rattus_norvegicus/Ensembl/RGSC3.4/Sequence/Bowtie2Index/genome SRR042499.fastq-o tophat_out : Output directory -p 10 : number of threads to use
Tophat2 is an aligner which takes account splice reads.
Tophat2 output
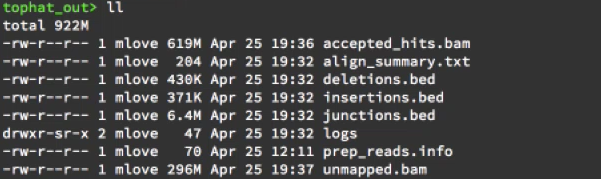
Read counting using Cufflinks
Cufflinks, estimates both the expression levels and the different RNA isoforms which are present in the sample. The main paper describing the Cufflinks method is Trapnell (2010) and the website for the Cufflinks software is:
http://cufflinks.cbcb.umd.edu/
After aligning the read, we can run cufflinks.
cufflinks -o cufflinks -p 10 --GTF-guide /path/to/Rattus_norvegicus/Ensembl/RGSC3.4/Annotation/Genes/genes.gtf \
tophat_out/accepted_hits.bam-o cufflinks: Output directory -p 10 : Number of thread to use –GTF-guide: indicate GTF file. This is the ensemble GTF file which contains all the information about the exons and transcripts and genes of rat.
And then finally, you give it the one or more files which have the aligned reads (accepted_hits.bam). This file contains also spliced reads.
Cufflinks will try to find the transcripts that are present and the level of the transcripts.
cufflinks output files are:
- genes.fpkm_tracking
- isoform.fpkm_tracking
- skipped.gtf
- transcriptd.gtf
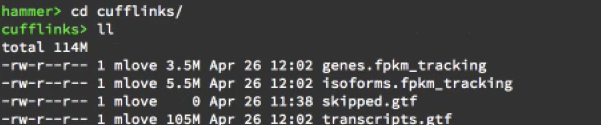
The first two are tracking files which gives some information about the genes. It gives here, in the last four columns, the measure of gene expression (FPKM, FPKM_conf_lo, FPKM_conf_hi, FPKM_status). FPKM is the number of fragments per kilobase per million mapped reads. So it’s normalized value of expression.
We are going to work with the transcripts GTF file. This contains information about all the transcripts which were found in the sample and the exons which belong to the transcript. You can see this file is a mix of known transcripts and in this case it will use the Ensembl ID (The ID of the GTF file you give it as a guide).
It also contains some transcripts which are given a new ID. this is a known transcript but the gene ID is annotated by Cufflinks. If there’s a new transcript it would also be given a cuff prefix. There is an example of both the gene is found by Cufflinks and a new transcript was found by Cufflinks.
Use IGV to visualize
I’m going to just show you what these look like in IGV. So I’ve downloaded the accepted hits.bam file and the transcripts .GTF file to my local directory. So I then use this line here to– it’s a grep line which removes any lines which match this. I wanted to remove lines which had FPKM 0.
grep -v ‘FPKM “0.0000000000”’ transcripts.gtf | less
And then I downloaded the chromosome 1 FASTA file and the rat GTF file so that I could look at these in IGV. So now here’s my IGV window where I’ve loaded in the coverage. And this line here is the ensemble GTF file and below it is the transcripts.gtf file from Cufflinks.
So they look very similar but we can zoom in and try to find some differences.
For visualizing:
ftp://ftp.ensembl.org/pub/release-69/fasta/rattus_norvegicus/dna/Rattus_norvegicus.RGSC3.4.69.dna.chromosome.1.fa.gz ftp://ftp.ensembl.org/pub/release-69/gtf/rattus_norvegicus/Rattus_norvegicus.RGSC3.4.69.gtf.gz
Licence: Licence References: https://github.com/genomicsclass
Show me some love with the like buttons below... Thank you and please don't forget to share and comment below!!
Montrez-moi un peu d'amour avec les like ci-dessous ... Merci et n'oubliez pas, s'il vous plaît, de partager et de commenter ci-dessous!
Recommended for You!
Recommended for you
This section contains the best data science and self-development resources to help you on your path.
Books - Data Science
Our Books
- Practical Guide to Cluster Analysis in R by A. Kassambara (Datanovia)
- Practical Guide To Principal Component Methods in R by A. Kassambara (Datanovia)
- Machine Learning Essentials: Practical Guide in R by A. Kassambara (Datanovia)
- R Graphics Essentials for Great Data Visualization by A. Kassambara (Datanovia)
- GGPlot2 Essentials for Great Data Visualization in R by A. Kassambara (Datanovia)
- Network Analysis and Visualization in R by A. Kassambara (Datanovia)
- Practical Statistics in R for Comparing Groups: Numerical Variables by A. Kassambara (Datanovia)
- Inter-Rater Reliability Essentials: Practical Guide in R by A. Kassambara (Datanovia)
Others
- R for Data Science: Import, Tidy, Transform, Visualize, and Model Data by Hadley Wickham & Garrett Grolemund
- Hands-On Machine Learning with Scikit-Learn, Keras, and TensorFlow: Concepts, Tools, and Techniques to Build Intelligent Systems by Aurelien Géron
- Practical Statistics for Data Scientists: 50 Essential Concepts by Peter Bruce & Andrew Bruce
- Hands-On Programming with R: Write Your Own Functions And Simulations by Garrett Grolemund & Hadley Wickham
- An Introduction to Statistical Learning: with Applications in R by Gareth James et al.
- Deep Learning with R by François Chollet & J.J. Allaire
- Deep Learning with Python by François Chollet
Click to follow us on Facebook :
Comment this article by clicking on "Discussion" button (top-right position of this page)







