RNA-Seq differential expression work flow using DESeq2
This analysis was performed using R (ver. 3.1.0).
Introduction
One of the aim of RNAseq data analysis is the detection of differentially expressed genes. The package DESeq2 provides methods to test for differential expression analysis.
This document presents an RNAseq differential expression workflow. We will start from the FASTQ files, align to the reference genome, prepare gene expression values as a count table by counting the sequenced fragments, perform differential gene expression analysis, and visually explore the results.
Input data
We will use publicly available data from the article by Felix Haglund et al., J Clin Endocrin Metab 2012. The purpose of the experiment was to investigate the role of the estrogen receptor in parathyroid tumors. The investigators derived primary cultures of parathyroid adenoma cells from 4 patients. These primary cultures were treated with diarylpropionitrile (DPN), an estrogen receptor beta agonist, or with 4-hydroxytamoxifen (OHT). RNA was extracted at 24 hours and 48 hours from cultures under treatment and control.
Part of the data from this experiment is provided in the Bioconductor data package parathyroidSE.
Aligning reads to a reference
What we get from the sequencing machine is a set of FASTQ files that contain the nucleotide sequence of each read and a quality score at each position. These reads must first be aligned to a reference genome or transcriptome. It is important to know if the sequencing experiment was single-end or paired-end, as the alignment software will require the user to specify both FASTQ files for a paired-end experiment. The output of this alignment step is commonly stored in a file format called BAM.
Here we use the TopHat2 spliced alignment software in combination with the Bowtie index available at the Illumina iGenomes.
For example, the paired-end RNA-Seq reads for the parathyroidSE package were aligned using TopHat2 with 8 threads, with the call:
tophat2 -o file_tophat_out -p 8 path/to/genome file_1.fastq file_2.fastq samtools sort -n file_tophat_out/accepted_hits.bam _sorted
The second line sorts the reads by name rather than by genomic position, which is necessary for counting paired-end reads within Bioconductor. This command uses the SAMtools software.
The BAM files for a number of sequencing runs can then be used to generate count matrices, as described in the following section.
Example BAM files
We will use BAM files from parathyroidSE package to demonstrate how a count table can be constructed from BAM files.
library( "parathyroidSE" )
#Find extdata
extDataDir <- system.file("extdata", package = "parathyroidSE", mustWork = TRUE)
list.files( extDataDir )## [1] "conversion.txt" "GSE37211_series_matrix.txt" "SRR479052.bam" "SRR479053.bam"
## [5] "SRR479054.bam"Typically, we have a table with experimental meta data for our samples. For these three files, it is as follows:
#Sample tables
sampleTable <- data.frame(
sampleName = c( "Ctrl_24h_1", "Ctrl_48h_1", "DPN_24h_1" ),
fileName = c( "SRR479052.bam", "SRR479053.bam", "SRR479054.bam" ),
treatment = c( "Control", "Control", "DPN" ),
time = c( "24h", "48h", "24h" ) )
#This is how the sample table should look like
sampleTable## sampleName fileName treatment time
## 1 Ctrl_24h_1 SRR479052.bam Control 24h
## 2 Ctrl_48h_1 SRR479053.bam Control 48h
## 3 DPN_24h_1 SRR479054.bam DPN 24hConstruct the full paths to the files we want to perform the counting operation on:
bamFiles <- file.path( extDataDir, sampleTable$fileName )
bamFiles## [1] "/Library/Frameworks/R.framework/Versions/3.1/Resources/library/parathyroidSE/extdata/SRR479052.bam"
## [2] "/Library/Frameworks/R.framework/Versions/3.1/Resources/library/parathyroidSE/extdata/SRR479053.bam"
## [3] "/Library/Frameworks/R.framework/Versions/3.1/Resources/library/parathyroidSE/extdata/SRR479054.bam"We can peek into one of the BAM files to see the naming style of the sequences (chromosomes). Here we use the BamFile function from the Rsamtools package.
library( "Rsamtools" )
seqinfo( BamFile( bamFiles[1] ) )## Seqinfo of length 25
## seqnames seqlengths isCircular genome
## 1 249250621
## 2 243199373
## 3 198022430
## 4 191154276
## 5 180915260
## ... ... ... ...
## 21 48129895
## 22 51304566
## X 155270560
## Y 59373566
## MT 16569 We want to make sure that these sequence names are the same style as that of the gene models we will obtain in the next section.
Counting reads in genes
To count how many read map to each gene, we need transcript annotation. Download the current GTF file with human gene annotation from Ensembl.
From this file, the function makeTranscriptDbFromGFF from the GenomicFeatures package constructs a database of all annotated transcripts.
For this lab you can use the truncated version of this file, called Homo_sapiens.GRCh37.75.subset.gtf.gz.
Preparing gene models from GFF files
library( "GenomicFeatures" )
#hse <- makeTranscriptDbFromGFF( "/path/to/your/genemodel_file.GTF", format="gtf" )
hse <- makeTranscriptDbFromGFF( "Homo_sapiens.GRCh37.75.subset.gtf", format="gtf" )
#Extract exons for each gene
exonsByGene <- exonsBy( hse, by="gene" )
exonsByGene## GRangesList of length 100:
## $ENSG00000000003
## GRanges with 13 ranges and 2 metadata columns:
## seqnames ranges strand | exon_id exon_name
## |
## [1] X [99883667, 99884983] - | 3038
## [2] X [99885756, 99885863] - | 3039
## [3] X [99887482, 99887565] - | 3040
## [4] X [99887538, 99887565] - | 3041
## [5] X [99888402, 99888536] - | 3042
## ... ... ... ... ... ... ...
## [9] X [99890555, 99890743] - | 3046
## [10] X [99891188, 99891686] - | 3047
## [11] X [99891605, 99891803] - | 3048
## [12] X [99891790, 99892101] - | 3049
## [13] X [99894942, 99894988] - | 3050
##
## ...
## <99 more elements>
## ---
## seqlengths:
## 7 12 2 6 16 4 3 1 17 8 19 X 11 9 20
## NA NA NA NA NA NA NA NA NA NA NA NA NA NA NA
- Note that gene models can also be prepared directly from BioMart :
library( "GenomicFeatures" )
# takes ~10 min
txdb <- makeTranscriptDbFromBiomart( biomart="ensembl",
dataset="hsapiens_gene_ensembl" )
exonsByGene <- exonsBy( txdb, by="gene" )Use saveDb() to only do this once. Use loadDb() to load the database next time.
library(TxDb.Hsapiens.UCSC.hg19.knownGene) is also an ready to go option for gene models.
- To convert from
chrXtoXand back:
seqlevelsStyle(gr) <- "NCBI"
seqlevelsStyle(gr) <- "UCSC"
Reads counting
The function summarizeOverlaps from the GenomicAlignments package will do this.
library( "GenomicAlignments" )
se <- summarizeOverlaps( exonsByGene, BamFileList( bamFiles ), mode="Union",
singleEnd=FALSE, ignore.strand=TRUE, fragments=TRUE )
We use the counting mode "Union", which indicates that those reads which overlap any portion of exactly one feature are counted. As this experiment produced paired-end reads, we specify singleEnd =FALSE. As protocol was not strand-specific, we specify ignore.strand = TRUE. fragments = TRUE indicates that we also want to count reads with unmapped pairs. This last argument is only for use with paired-end experiments.
Details on how to read from the BAM files can be specified using the BamFileList function. For example, to control the memory, we could have specified that batches of 2 000 000 reads should be read at a time:
BamFileList( bamFiles, yieldSize = 2000000 )
SummarizedExperiment object : Output of counting
We investigate the resulting SummarizedExperiment class by looking at the counts in the assay slot, the phenotypic data about the samples in colData slot (in this case an empty DataFrame), and the data about the genes in the rowData slot.
Figure 1 explains the basic structure of the SummarizedExperiment class.
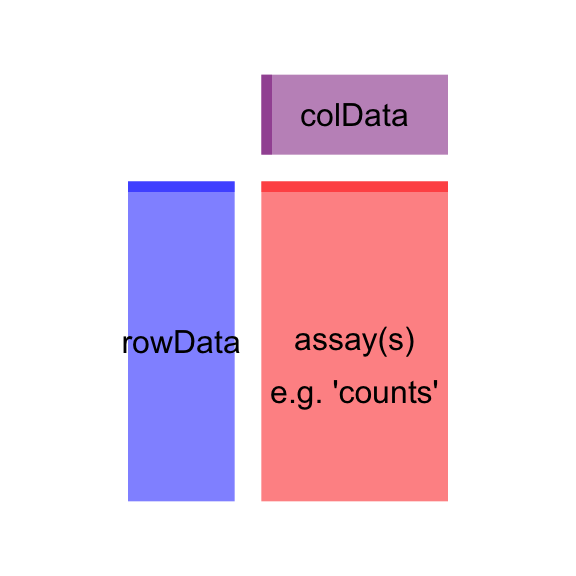
# SummarizedExperiment
se## class: SummarizedExperiment
## dim: 100 3
## exptData(0):
## assays(1): counts
## rownames(100): ENSG00000000003 ENSG00000000005 ... ENSG00000005469 ENSG00000005471
## rowData metadata column names(0):
## colnames(3): SRR479052.bam SRR479053.bam SRR479054.bam
## colData names(0):# Count data
head( assay(se) )## SRR479052.bam SRR479053.bam SRR479054.bam
## ENSG00000000003 0 0 1
## ENSG00000000005 0 0 0
## ENSG00000000419 0 0 0
## ENSG00000000457 0 1 1
## ENSG00000000460 0 0 0
## ENSG00000000938 0 0 0#Column sum
colSums( assay(se) )## SRR479052.bam SRR479053.bam SRR479054.bam
## 31 21 30#Phenotypic data
colData(se)## DataFrame with 3 rows and 0 columns#Data about genes
rowData(se)## GRangesList of length 100:
## $ENSG00000000003
## GRanges with 13 ranges and 2 metadata columns:
## seqnames ranges strand | exon_id exon_name
## |
## [1] X [99883667, 99884983] - | 3038
## [2] X [99885756, 99885863] - | 3039
## [3] X [99887482, 99887565] - | 3040
## [4] X [99887538, 99887565] - | 3041
## [5] X [99888402, 99888536] - | 3042
## ... ... ... ... ... ... ...
## [9] X [99890555, 99890743] - | 3046
## [10] X [99891188, 99891686] - | 3047
## [11] X [99891605, 99891803] - | 3048
## [12] X [99891790, 99892101] - | 3049
## [13] X [99894942, 99894988] - | 3050
##
## ...
## <99 more elements>
## ---
## seqlengths:
## 7 12 2 6 16 4 3 1 17 8 19 X 11 9 20
## NA NA NA NA NA NA NA NA NA NA NA NA NA NA NA Note that the rowData slot is a GRangesList, which contains all the information about the exons for each gene, i.e., for each row of the count table.
The colData slot, so far empty, should contain all the meta data. We hence assign our sample table to it:
#Column data
colData(se) <- DataFrame( sampleTable )We can extract columns from the colData using the $ operator, and we can omit the colData to avoid extra keystrokes.
colData(se)$treatment## [1] Control Control DPN
## Levels: Control DPNse$treatment## [1] Control Control DPN
## Levels: Control DPNWe can also use the sampleName table to name the columns of our data matrix:
colnames(se) <- sampleTable$sampleName
head( assay(se) )## Ctrl_24h_1 Ctrl_48h_1 DPN_24h_1
## ENSG00000000003 0 0 1
## ENSG00000000005 0 0 0
## ENSG00000000419 0 0 0
## ENSG00000000457 0 1 1
## ENSG00000000460 0 0 0
## ENSG00000000938 0 0 0This SummarizedExperiment object se is then all we need to start our analysis.The DESeqDataSet, column metadata, and the design formula
The data object class in DESeq2 is the DESeqDataSet, which is built on top of the SummarizedExperiment class. One main differences is that the assay slot is instead accessed using the count accessor, and the values in this matrix must be non-negative integers.
A second difference is that the DESeqDataSet has an associated “design formula”. The design formula tells which variables in the column metadata table colData specify the experimental design and how these factors should be used in the analysis.
The simplest design formula for differential expression would be ~ condition, where condition is a column in colData(dds) which specifies which of two (or more groups) the samples belong to.
For the parathyroid experiment, we will specify ~ patient + treatment, which means that we want to test for the effect of treatment (the last factor), controlling for the effect of patient (the first factor).
We now use R’s data command to load a prepared SummarizedExperiment that was generated from the publicly available sequencing data files associated with the Haglund et al. paper, described on page 1. The steps we used to produce this object were equivalent to those you worked through in the previous Section, except that we used the complete set of samples and all reads.
#Run data
data( "parathyroidGenesSE" )
se <- parathyroidGenesSE
colnames(se) <- se$run
# Check column data
colData(se)[1:5,1:4]## DataFrame with 5 rows and 4 columns
## run experiment patient treatment
##
## SRR479052 SRR479052 SRX140503 1 Control
## SRR479053 SRR479053 SRX140504 1 Control
## SRR479054 SRR479054 SRX140505 1 DPN
## SRR479055 SRR479055 SRX140506 1 DPN
## SRR479056 SRR479056 SRX140507 1 OHT Here we see that this object already contains an informative colData slot.
When you work with your own data, you will have to add the pertinent sample / phenotypic information for the experiment at this stage. We highly recommend keeping this information in a comma-separated value (CSV) or tab-separated value (TSV) file, which can be exported from an Excel spreadsheet, and the assign this to the colData slot, as shown in the previous section.
Make sure that the order of rows in your column data table matches the order of columns in the assay data slot.A bonus about the workflow we have shown above is that information about the gene models we used is included without extra effort. The str R function is used to compactly display the structure of the data in the list.
str( metadata( rowData(se) ) )## List of 1
## $ genomeInfo:List of 20
## ..$ Db type : chr "TranscriptDb"
## ..$ Supporting package : chr "GenomicFeatures"
## ..$ Data source : chr "BioMart"
## ..$ Organism : chr "Homo sapiens"
## ..$ Resource URL : chr "www.biomart.org:80"
## ..$ BioMart database : chr "ensembl"
## ..$ BioMart database version : chr "ENSEMBL GENES 72 (SANGER UK)"
## ..$ BioMart dataset : chr "hsapiens_gene_ensembl"
## ..$ BioMart dataset description : chr "Homo sapiens genes (GRCh37.p11)"
## ..$ BioMart dataset version : chr "GRCh37.p11"
## ..$ Full dataset : chr "yes"
## ..$ miRBase build ID : chr NA
## ..$ transcript_nrow : chr "213140"
## ..$ exon_nrow : chr "737783"
## ..$ cds_nrow : chr "531154"
## ..$ Db created by : chr "GenomicFeatures package from Bioconductor"
## ..$ Creation time : chr "2013-07-30 17:30:25 +0200 (Tue, 30 Jul 2013)"
## ..$ GenomicFeatures version at creation time: chr "1.13.21"
## ..$ RSQLite version at creation time : chr "0.11.4"
## ..$ DBSCHEMAVERSION : chr "1.0"Once we have our fully annotated SummerizedExperiment object, we can construct a DESeqDataSet object from it, which will then form the staring point of the actual DESeq2 package
library( "DESeq2" )
ddsFull <- DESeqDataSet( se, design = ~ patient + treatment )Note that there are two alternative functions, DESeqDataSetFromMatrix and DESeqDataSetFromHTSeq, which allow you to get started in case you have your data not in the form of a SummarizedExperiment object, but either as a simple matrix of count values or as output files from the htseq-count script from the HTSeq Python package.
Collapsing technical replicates
There are a number of samples which were sequenced in multiple runs. For example, sample SRS308873 was sequenced twice.
#as.data.frame forces R to show us the full list
head(as.data.frame( colData( ddsFull )[ ,c("sample","patient","treatment","time") ] ), 12)## sample patient treatment time
## SRR479052 SRS308865 1 Control 24h
## SRR479053 SRS308866 1 Control 48h
## SRR479054 SRS308867 1 DPN 24h
## SRR479055 SRS308868 1 DPN 48h
## SRR479056 SRS308869 1 OHT 24h
## SRR479057 SRS308870 1 OHT 48h
## SRR479058 SRS308871 2 Control 24h
## SRR479059 SRS308872 2 Control 48h
## SRR479060 SRS308873 2 DPN 24h
## SRR479061 SRS308873 2 DPN 24h
## SRR479062 SRS308874 2 DPN 48h
## SRR479063 SRS308875 2 OHT 24hA convenience function has been implemented to collapse, which can take an object, either SummarizedExperiment or DESeqDataSet, and a grouping factor, in this case the sample name, and return the object with the counts summed up for each unique sample. Optionally, we can provide a third argument, run, which can be used to paste together the names of the runs which were collapsed to create the new object.
Note that dds$variable is equivalent to colData(dds)$variable.ddsCollapsed <- collapseReplicates( ddsFull,
groupby = ddsFull$sample,
run = ddsFull$run )
head( as.data.frame( colData(ddsCollapsed)[ ,c("sample","runsCollapsed") ] ), 12 )## sample runsCollapsed
## SRS308865 SRS308865 SRR479052
## SRS308866 SRS308866 SRR479053
## SRS308867 SRS308867 SRR479054
## SRS308868 SRS308868 SRR479055
## SRS308869 SRS308869 SRR479056
## SRS308870 SRS308870 SRR479057
## SRS308871 SRS308871 SRR479058
## SRS308872 SRS308872 SRR479059
## SRS308873 SRS308873 SRR479060,SRR479061
## SRS308874 SRS308874 SRR479062
## SRS308875 SRS308875 SRR479063,SRR479064
## SRS308876 SRS308876 SRR479065We can confirm that the counts for the new object are equal to the summed up counts of the columns that had the same value for the grouping factor:
original <- rowSums( counts(ddsFull)[ , ddsFull$sample == "SRS308873" ] )
all( original == counts(ddsCollapsed)[ ,"SRS308873" ] )## [1] TRUERunning the DESeq2 pipeline
Here we will analyze a subset of the samples, namely those taken after 48 hours, with either control, DPN or OHT treatment, taking into account the multifactor design.
Preparing the data object for the analysis of interest
First we subset the relevant columns from the full dataset:
dds <- ddsCollapsed[ , ddsCollapsed$time == "48h" ]Sometimes it is necessary to drop levels of the factors, in case that all the samples for one or more levels of a factor in the design have been removed. If time were included in the design formula, the following code could be used to take care of dropped levels in this column.
dds$time <- droplevels( dds$time )It will be convenient to make sure that Control is the first level in the treatment factor, so that the default log2 fold changes are calculated as treatment over control and not the other way around. The function relevel achieves this:
dds$treatment <- relevel( dds$treatment, "Control" )A quick check whether we now have the right samples:
as.data.frame( colData(dds) )## run experiment patient treatment time submission study sample runsCollapsed
## SRS308866 SRR479053 SRX140504 1 Control 48h SRA051611 SRP012167 SRS308866 SRR479053
## SRS308868 SRR479055 SRX140506 1 DPN 48h SRA051611 SRP012167 SRS308868 SRR479055
## SRS308870 SRR479057 SRX140508 1 OHT 48h SRA051611 SRP012167 SRS308870 SRR479057
## SRS308872 SRR479059 SRX140510 2 Control 48h SRA051611 SRP012167 SRS308872 SRR479059
## SRS308874 SRR479062 SRX140512 2 DPN 48h SRA051611 SRP012167 SRS308874 SRR479062
## SRS308876 SRR479065 SRX140514 2 OHT 48h SRA051611 SRP012167 SRS308876 SRR479065
## SRS308878 SRR479067 SRX140516 3 Control 48h SRA051611 SRP012167 SRS308878 SRR479067
## SRS308880 SRR479069 SRX140518 3 DPN 48h SRA051611 SRP012167 SRS308880 SRR479069
## SRS308882 SRR479071 SRX140520 3 OHT 48h SRA051611 SRP012167 SRS308882 SRR479071
## SRS308883 SRR479072 SRX140521 4 Control 48h SRA051611 SRP012167 SRS308883 SRR479072
## SRS308885 SRR479074 SRX140523 4 DPN 48h SRA051611 SRP012167 SRS308885 SRR479074,SRR479075
## SRS308887 SRR479077 SRX140525 4 OHT 48h SRA051611 SRP012167 SRS308887 SRR479077,SRR479078In order to speed up some annotation steps below, it makes sense to remove genes which have zero counts for all samples. This can be done by simply indexing the dds object:
dds <- dds[ rowSums( counts(dds) ) > 0 , ]Running the pipeline
Let’s recall what design we have specified:
#design
design(dds)## ~patient + treatment#Run DESeq : Modeling counts with patient and treatment effects
dds <- DESeq(dds)This function will print out a message for the various steps it performs. Briefly these are: the estimation of size factors (which control for differences in the library size of the sequencing experiments), the estimation of dispersion for each gene, and fitting a generalized linear model.A DESeqDataSet is returned which contains all the fitted information within it, and the following section describes how to extract out results tables of interest from this object.
Inspecting the results table
Calling results without any arguments will extract the estimated log2 fold changes and p values for the last variable in the design formula. If there are more than 2 levels for this variable – as is the case in this analysis – results will extract the results table for a comparison of the last level over the first level. The following section describes how to extract other comparisons.
res <- results( dds )
res## log2 fold change (MAP): treatment OHT vs Control
## Wald test p-value: treatment OHT vs Control
## DataFrame with 32082 rows and 6 columns
## baseMean log2FoldChange lfcSE stat pvalue padj
##
## ENSG00000000003 613.8196 -0.044799 0.08787 -0.50983 0.610173 0.9837
## ENSG00000000005 0.5502 -0.568326 1.08754 -0.52258 0.601268 NA
## ENSG00000000419 304.0482 0.116118 0.09623 1.20668 0.227555 NA
## ENSG00000000457 183.5157 0.007441 0.12314 0.06042 0.951819 NA
## ENSG00000000460 207.4336 0.470836 0.14493 3.24867 0.001159 NA
## ... ... ... ... ... ... ...
## ENSG00000271699 0.17125 -1.09753 0.9097 -1.20646 0.22764 NA
## ENSG00000271704 0.10233 -0.01414 0.7157 -0.01975 0.98424 NA
## ENSG00000271707 9.21112 -0.84445 0.4832 -1.74774 0.08051 NA
## ENSG00000271709 0.08257 -0.01108 0.6782 -0.01633 0.98697 NA
## ENSG00000271711 0.74835 -0.95188 1.0655 -0.89335 0.37167 NA As res is a DataFrame object, it carries metadata with information on the meaning of the columns:
mcols(res, use.names=TRUE)## DataFrame with 6 rows and 2 columns
## type description
##
## baseMean intermediate the base mean over all rows
## log2FoldChange results log2 fold change (MAP): treatment OHT vs Control
## lfcSE results standard error: treatment OHT vs Control
## stat results Wald statistic: treatment OHT vs Control
## pvalue results Wald test p-value: treatment OHT vs Control
## padj results BH adjusted p-values The first column, baseMean, is a just the average of the normalized count values, dividing by size factors, taken over all samples. The remaining four columns refer to a specific contrast, namely the comparison of the levels DPN versus Control of the factor variable treatment.
See the help page for results (by typing ?results) for information on how to obtain other contrasts.
The column log2FoldChange is the effect size estimate. It tells us how much the gene’s expression seems to have changed due to treatment with DPN in comparison to control. This value is reported on a logarithmic scale to base 2: for example, a log2 fold change of 1.5 means that the gene’s expression is increased by a multiplicative factor of 21.5 ≈ 2.82.
Of course, this estimate has an uncertainty associated with it, which is available in the column lfcSE, the standard error estimate for the log2 fold change estimate. The column p value indicates wether the observed difference between treatment and control is significantly different.
We note that a subset of the p values in res are NA (“notavailable”). This is DESeq’s way of reporting that all counts for this gene were zero, and hence not test was applied. In addition, p values can be assigned NA if the gene was excluded from analysis because it contained an extreme count outlier. For more information, see the outlier detection section of the advanced vignette.
We can examine the counts and normalized counts for the gene with the smallest p value:
# SmallestPvalue
idx <- which.min(res$pvalue)
counts(dds)[idx, ]## SRS308866 SRS308868 SRS308870 SRS308872 SRS308874 SRS308876 SRS308878 SRS308880 SRS308882 SRS308883 SRS308885 SRS308887
## 95 62 30 24 13 21 276 357 112 388 404 296#Normalization
counts(dds, normalized=TRUE)[ idx, ]## SRS308866 SRS308868 SRS308870 SRS308872 SRS308874 SRS308876 SRS308878 SRS308880 SRS308882 SRS308883 SRS308885 SRS308887
## 86.81 61.43 37.42 35.25 15.96 25.85 314.96 201.94 136.92 461.46 269.09 172.00Other comparisons
The results for a comparison of any two levels of a variable can be extracted using the contrast argument to results. The user should specify three values: The name of the variable, the name of the level in the numerator, and the name of the level in the denominator.
Here we extract results for the log2 of the fold change of DPN/Control:
#Save result table
resOHT <- res
#Other comparisons
res <- results( dds, contrast = c("treatment", "DPN", "Control") )
res## log2 fold change (MAP): treatment DPN vs Control
## Wald test p-value: treatment DPN vs Control
## DataFrame with 32082 rows and 6 columns
## baseMean log2FoldChange lfcSE stat pvalue padj
##
## ENSG00000000003 613.8196 -0.01722 0.08665 -0.19868 0.84252 0.9775
## ENSG00000000005 0.5502 -0.10344 1.09363 -0.09458 0.92465 NA
## ENSG00000000419 304.0482 -0.01695 0.09517 -0.17807 0.85867 0.9811
## ENSG00000000457 183.5157 -0.09654 0.12138 -0.79533 0.42642 0.8945
## ENSG00000000460 207.4336 0.35004 0.14375 2.43497 0.01489 0.2762
## ... ... ... ... ... ... ...
## ENSG00000271699 0.17125 -1.1391 0.9251 -1.2313 0.2182 NA
## ENSG00000271704 0.10233 0.5925 0.7547 0.7851 0.4324 NA
## ENSG00000271707 9.21112 -0.5099 0.4579 -1.1136 0.2655 NA
## ENSG00000271709 0.08257 0.5156 0.7112 0.7250 0.4685 NA
## ENSG00000271711 0.74835 -0.6649 1.0511 -0.6326 0.5270 NA Adding gene names
Our result table only uses Ensembl gene IDs, but gene names may be more informative. Bioconductor’s annotation packages help with mapping various ID schemes to each other.
We load the annotation package org.Hs.eg.db:
library( "org.Hs.eg.db" )This is the organism annotation package (“org”“) for Homo sapiens (”Hs“”), organized as an AnnotationDbi package (“db”“), using Entrez Gene IDs (”eg“) as primary key.
To get a list of all available key types, use
columns(org.Hs.eg.db)## [1] "ENTREZID" "PFAM" "IPI" "PROSITE" "ACCNUM" "ALIAS" "CHR"
## [8] "CHRLOC" "CHRLOCEND" "ENZYME" "MAP" "PATH" "PMID" "REFSEQ"
## [15] "SYMBOL" "UNIGENE" "ENSEMBL" "ENSEMBLPROT" "ENSEMBLTRANS" "GENENAME" "UNIPROT"
## [22] "GO" "EVIDENCE" "ONTOLOGY" "GOALL" "EVIDENCEALL" "ONTOLOGYALL" "OMIM"
## [29] "UCSCKG"Converting IDs with the native functions from the AnnotationDbi package is currently a bit cumbersome, so we provide the following convenience function (without explaining how exactly it works):
#ids = list of IDS
#fromKey = key type; toKey = key type we want to convert to
#db = the AnnotationDb object to use.
#ifMultiple = the argument specifies what to do if one source ID maps to several target IDs:
#should the function return an NA or simply the first of the multiple IDs?
convertIDs <- function( ids, fromKey, toKey, db, ifMultiple=c( "putNA", "useFirst" ) ) {
stopifnot( inherits( db, "AnnotationDb" ) )
ifMultiple <- match.arg( ifMultiple )
suppressWarnings( selRes <- AnnotationDbi::select(
db, keys=ids, keytype=fromKey, columns=c(fromKey,toKey) ) )
if( ifMultiple == "putNA" ) {
duplicatedIds <- selRes[ duplicated( selRes[,1] ), 1 ]
selRes <- selRes[ ! selRes[,1] %in% duplicatedIds, ] }
return( selRes[ match( ids, selRes[,1] ), 2 ] )
}To convert the Ensembl IDs in the rownames of res to gene symbols and add them as a new column, we use:
res$hgnc_symbol <- convertIDs( row.names(res), "ENSEMBL", "SYMBOL", org.Hs.eg.db )
res$entrezid <- convertIDs( row.names(res), "ENSEMBL", "ENTREZID", org.Hs.eg.db )
head(res, 4)## log2 fold change (MAP): treatment DPN vs Control
## Wald test p-value: treatment DPN vs Control
## DataFrame with 4 rows and 8 columns
## baseMean log2FoldChange lfcSE stat pvalue padj hgnc_symbol entrezid
##
## ENSG00000000003 613.8196 -0.01722 0.08665 -0.19868 0.8425 0.9775 TSPAN6 7105
## ENSG00000000005 0.5502 -0.10344 1.09363 -0.09458 0.9246 NA TNMD 64102
## ENSG00000000419 304.0482 -0.01695 0.09517 -0.17807 0.8587 0.9811 DPM1 8813
## ENSG00000000457 183.5157 -0.09654 0.12138 -0.79533 0.4264 0.8945 SCYL3 57147 This conversion can be done from Ensembl BioMart web site.Further points
Multiple testing
DESeq2 uses the so-called Benjamini-Hochberg (BH) adjustment for multiple testing problem; in brief, this method calculates for each gene an adjusted p value which answers the following question: if one called significant all genes with a p value less than or equal to this gene’s p value threshold, what would be the fraction of false positives (the false discovery rate, FDR) among them (in the sense of the calculation outlined above)? These values, called the BH-adjusted p values, are given in the column padj of the results object.
Hence, if we consider a fraction of 10% false positives acceptable, we can consider all genes with an adjusted p value below 10%=0.1 as significant. How many such genes are there?
sum( res$padj < 0.1, na.rm=TRUE )## [1] 249We subset the results table to these genes and then sort it by the log2 fold change estimate to get the significant genes with the strongest down-regulation:
resSig <- subset(res, res$padj < 0.1 )
head( resSig[ order( resSig$log2FoldChange ), ], 4)## log2 fold change (MAP): treatment DPN vs Control
## Wald test p-value: treatment DPN vs Control
## DataFrame with 4 rows and 8 columns
## baseMean log2FoldChange lfcSE stat pvalue padj hgnc_symbol entrezid
##
## ENSG00000163631 233.3 -0.9307 0.2842 -3.275 1.058e-03 0.05817 ALB 213
## ENSG00000119946 151.6 -0.6903 0.1566 -4.408 1.041e-05 0.00246 CNNM1 26507
## ENSG00000041982 1377.0 -0.6859 0.1845 -3.717 2.018e-04 0.02045 TNC 3371
## ENSG00000155111 530.9 -0.6756 0.2109 -3.203 1.359e-03 0.06850 CDK19 23097 and with the strongest upregulation :
head( resSig[ order( -resSig$log2FoldChange ), ], 4)## log2 fold change (MAP): treatment DPN vs Control
## Wald test p-value: treatment DPN vs Control
## DataFrame with 4 rows and 8 columns
## baseMean log2FoldChange lfcSE stat pvalue padj hgnc_symbol entrezid
##
## ENSG00000092621 558.5 0.9002 0.1258 7.158 8.191e-13 2.628e-09 PHGDH 26227
## ENSG00000101255 255.1 0.8860 0.1694 5.229 1.706e-07 2.346e-04 TRIB3 57761
## ENSG00000103257 168.5 0.8259 0.1644 5.024 5.064e-07 3.249e-04 SLC7A5 8140
## ENSG00000156414 142.8 0.7582 0.1661 4.564 5.025e-06 1.547e-03 TDRD9 122402 Diagnostic plot
MA plot
A so-called MA plot provides a useful overview for an experiment with a two-group comparison:
plotMA( res, ylim = c(-3, 3) )
The MA-plot represents each gene with a dot. The x axis is the average expression over all samples, the y axis the log2 fold change of normalized counts (i.e the average of counts normalized by size factor) between treatment and control. Genes with an adjusted p value below a threshold (here 0.1, the default) are shown in red.
This plot demonstrates that only genes with a large average normalized count contain sufficient information to yield a significant call.
Dispersion plot
Also note DESeq2 shrinkage estimation of log fold changes (LFCs): When count values are too low to allow an accurate estimate of the LFC, the value is “shrunken”" towards zero to avoid that these values, which otherwise would frequently be unrealistically large, dominate the top-ranked log fold change. Whether a gene is called significant depends not only on its LFC but also on its within-group variability, which DESeq2 quantifies as the dispersion. For strongly expressed genes, the dispersion can be understood as a squared coefficient of variation: a dispersion value of 0.01 means that the gene’s expression tends to differ by typically $\sqrt{0.01}=10\%$ between samples of the same treatment group. For weak genes, the Poisson noise is an additional source of noise, which is added to the dispersion. The function plotDispEsts visualizes DESeq2’s dispersion estimates:
plotDispEsts( dds, ylim = c(1e-6, 1e1) )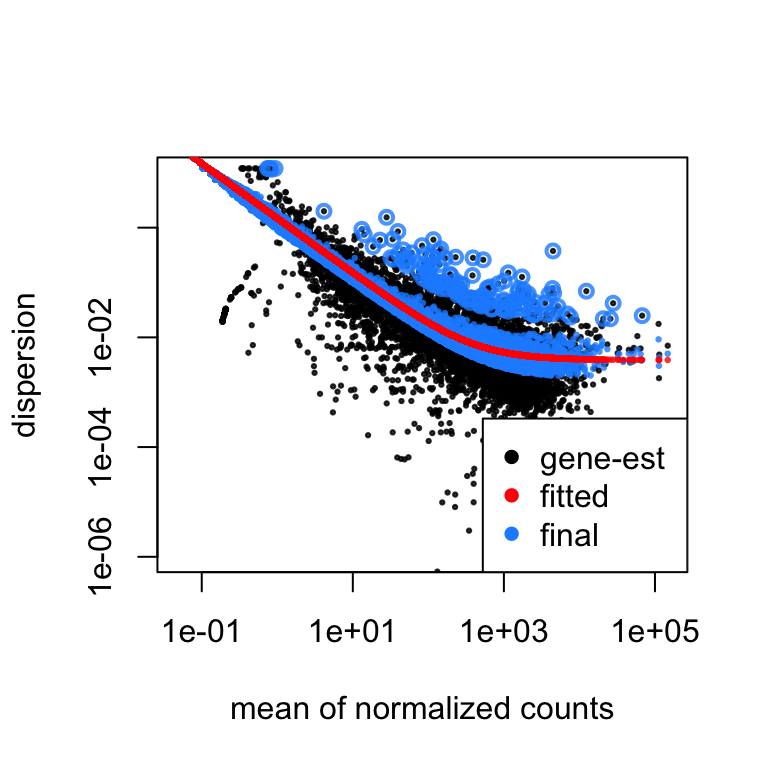
The black points are the dispersion estimates for each gene as obtained by considering the information from each gene separately. Unless one has many samples, these values fluctuate strongly around their true values. Therefore, we fit the red trend line, which shows the dispersions’ dependence on the mean, and then shrink each gene’s estimate towards the red line to obtain the final estimates (blue points) that are then used in the hypothesis test. The blue circles above the main “cloud”" of points are genes which have high gene-wise dispersion estimates which are labelled as dispersion outliers. These estimates are therefore not shrunk toward the fitted trend line.
Histogram of the p-value
hist( res$pvalue, breaks=20, col="grey" )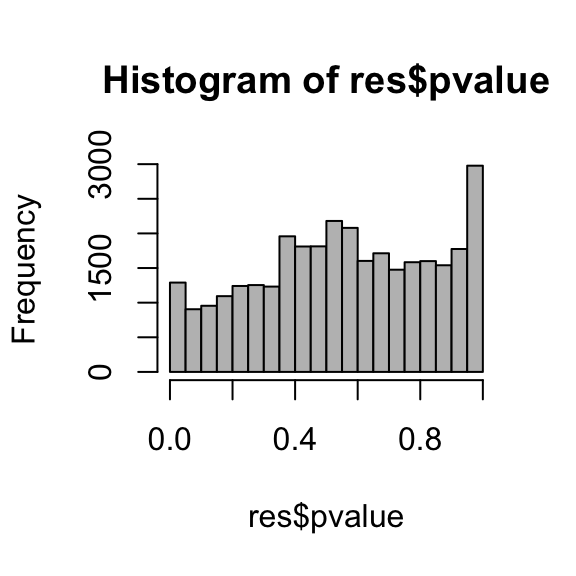
Independent filtering
The MA plot highlights an important property of RNA-Seq data. For weakly expressed genes, we have no chance of seeing differential expression, because the low read counts suffer from so high Poisson noise that any biological effect is drowned in the uncertainties from the read counting. We can also show this by examining the ratio of small p values (say, less than, 0.01) for genes binned by mean normalized count:
# create bins using the quantile function
qs <- c( 0, quantile( res$baseMean[res$baseMean > 0], 0:7/7 ) )
# "cut" the genes into the bins
bins <- cut( res$baseMean, qs )
# rename the levels of the bins using the middle point
levels(bins) <- paste0("~",round(.5*qs[-1] + .5*qs[-length(qs)]))
# calculate the ratio of p values less than .01 for each bin
ratios <- tapply( res$pvalue, bins, function(p) mean( p < .01, na.rm=TRUE ) )
# plot these ratios
barplot(ratios, xlab="mean normalized count", ylab="ratio of small p values")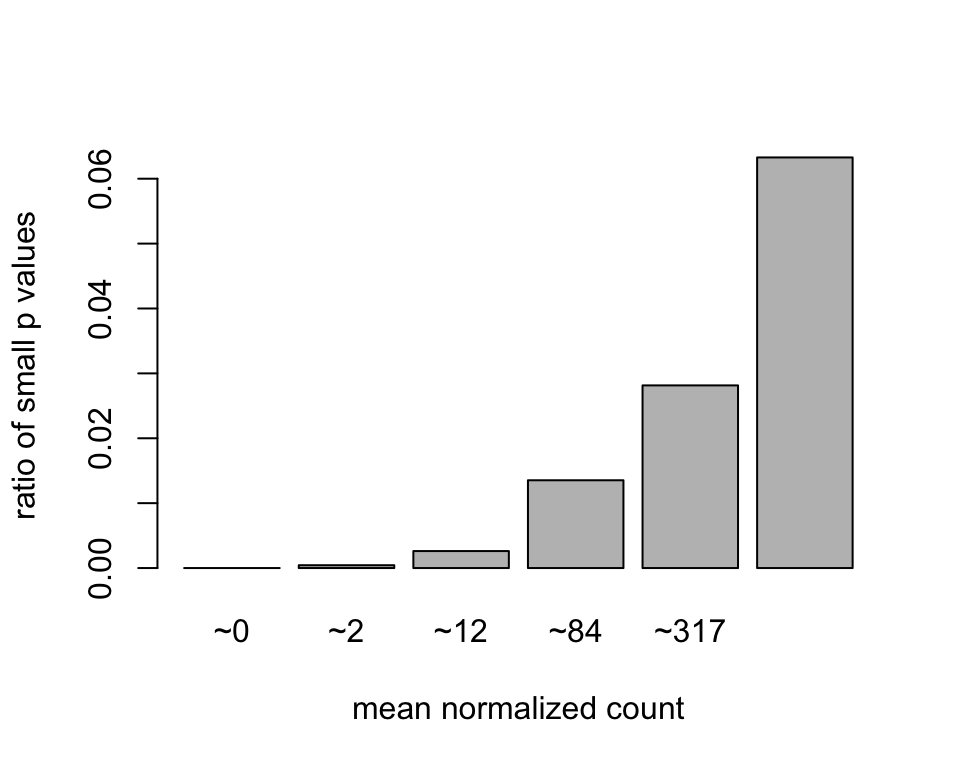
At first sight, there may seem to be little benefit in filtering out these genes. After all, the test found them to be non-significant anyway. However, these genes have an influence on the multiple testing adjustment, whose performance improves if such genes are removed. By removing the weakly-expressed genes from the input to the FDR procedure, we can find more genes to be significant among those which we keep, and so improved the power of our test. This approach is known as independent filtering.
The DESeq software automatically performs independent filtering which maximizes the number of genes which will have adjusted p value less than a critical value (by default, alpha is set to 0.1). This automatic independent filtering is performed by, and can be controlled by, the results function. We can observe how the number of rejections changes for various cutoffs based on mean normalized count. The following optimal threshold and table of possible values is stored as an attribute of the results object.
attr(res,"filterThreshold")## 70%
## 126.9plot(attr(res,"filterNumRej"),type="b",
xlab="quantiles of 'baseMean'",
ylab="number of rejections")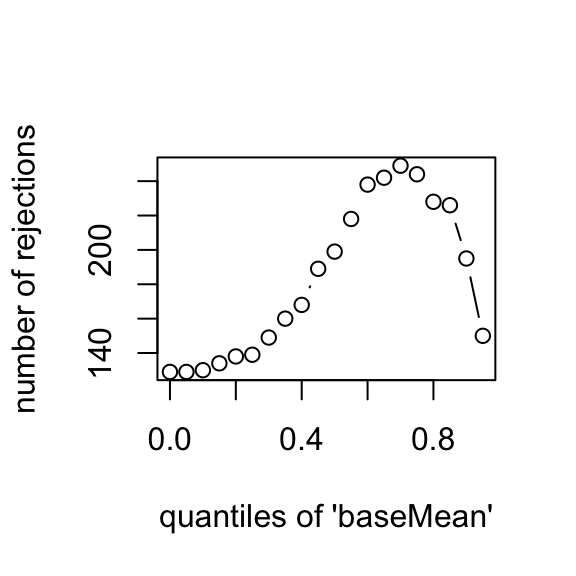
The term independent highlights an important caveat. Such filtering is permissible only if the filter criterion is independent of the actual test statistic. Otherwise, the filtering would invalidate the test and consequently the assumptions of the BH procedure. This is why we filtered on the average over all samples: this filter is blind to the assignment of samples to the treatment and control group and hence independent.
Exporting results
You can easily save the results table in a CSV file, which you can then load with a spreadsheet program such as Excel:
res[1:2,]
write.csv( as.data.frame(res), file="results.csv" )Gene-set enrichment analysis
Do the genes with a strong up- or down-regulation have something in common? We perform next a gene-set enrichment analysis (GSEA) to examine this question.
We here present a relatively simplistic approach, to demonstrate the basic ideas, but note that a more careful treatment will be needed for more definitive results.
We use the gene sets in the Reactome database:
#source("http://bioconductor.org/biocLite.R")
#biocLite("reactome.db")
library( "reactome.db" )This database works with Entrez IDs, so we will need the entrezid column that we added earlier to the res object.
First, we subset the results table, res, to only those genes for which the Reactome database has data (i.e, whose Entrez ID we find in the respective key column of reactome.db and for which the DESeq2 test gave an adjusted p value that was not NA.
res2 <- res[ res$entrezid %in% keys( reactome.db, "ENTREZID" ) & !is.na( res$padj ) , ]
head(res2)## log2 fold change (MAP): treatment DPN vs Control
## Wald test p-value: treatment DPN vs Control
## DataFrame with 6 rows and 8 columns
## baseMean log2FoldChange lfcSE stat pvalue padj hgnc_symbol entrezid
##
## ENSG00000000419 304.0 -0.016946 0.09517 -0.17807 0.8587 0.9811 DPM1 8813
## ENSG00000001084 312.0 0.039811 0.10228 0.38923 0.6971 0.9540 GCLC 2729
## ENSG00000001167 398.8 -0.056003 0.09711 -0.57669 0.5641 0.9357 NFYA 4800
## ENSG00000001630 463.5 0.070002 0.09676 0.72348 0.4694 0.9123 CYP51A1 1595
## ENSG00000002822 169.5 -0.001926 0.13784 -0.01397 0.9889 0.9982 MAD1L1 8379
## ENSG00000003056 995.8 0.007716 0.06785 0.11373 0.9095 0.9859 M6PR 4074 Using select, a function from AnnotationDbi for querying database objects, we get a table with the mapping from Entrez IDs to Reactome Path IDs :
reactomeTable <- AnnotationDbi::select( reactome.db,
keys=as.character(res2$entrezid), keytype="ENTREZID",
columns=c("ENTREZID","REACTOMEID") )
head(reactomeTable)## ENTREZID REACTOMEID
## 1 8813 162699
## 2 8813 163125
## 3 8813 392499
## 4 8813 446193
## 5 8813 446203
## 6 8813 446219The next code chunk transforms this table into an incidence matrix. This is a Boolean matrix with one row for each Reactome Path and one column for each unique gene in res2, which tells us which genes are members of which Reactome Paths.
#problem with this code is that res2$entrez are not unique
#ft2mat <- function(x, y) {
# ux = unique(x)
# uy = unique(y)
# im = matrix(FALSE, nrow=length(ux), ncol=length(uy), dimnames=list(ux, uy))
# im[ cbind(x, y) ] = TRUE
# return(im)
#}
#incm <- with(reactomeTable, ft2mat(REACTOMEID, ENTREZID))incm <- do.call( rbind, with(reactomeTable, tapply(
ENTREZID, factor(REACTOMEID), function(x) res2$entrezid %in% x ) ))
colnames(incm) <- res2$entrez
str(incm)## logi [1:1458, 1:3400] FALSE FALSE FALSE FALSE FALSE FALSE ...
## - attr(*, "dimnames")=List of 2
## ..$ : chr [1:1458] "1059683" "109581" "109582" "109606" ...
## ..$ : chr [1:3400] "8813" "2729" "4800" "1595" ...We remove all rows corresponding to Reactome Paths with less than 20 or more than 80 assigned genes.
within <- function(x, lower, upper) (x>=lower & x<=upper)
incm <- incm[ within(rowSums(incm), lower=20, upper=80), ]To test whether the genes in a Reactome Path behave in a special way in our experiment, we calculate a number of statistics, including a t-statistic to see whether the average of the genes’ log2 fold change values in the gene set is different from zero. To facilitate the computations, we define a little helper function:
testCategory <- function( reactomeID ) {
isMember <- incm[ reactomeID, ]
data.frame(
reactomeID = reactomeID,
numGenes = sum( isMember ),
avgLFC = mean( res2$log2FoldChange[isMember] ),
sdLFC = sd( res2$log2FoldChange[isMember] ),
zValue = mean( res2$log2FoldChange[isMember] ) /sd( res2$log2FoldChange[isMember] ),
strength = sum( res2$log2FoldChange[isMember] ) / sqrt(sum(isMember)),
pvalue = t.test( res2$log2FoldChange[ isMember ] )$p.value,
reactomeName = reactomePATHID2NAME[[reactomeID]],
stringsAsFactors = FALSE ) }The function can be called with a Reactome Path ID:
testCategory("109606")## reactomeID numGenes avgLFC sdLFC zValue strength pvalue reactomeName
## 1 109606 31 -0.02215 0.07769 -0.2852 -0.1234 0.1228 Homo sapiens: Intrinsic Pathway for ApoptosisAs you can see the function not only performs the t test and returns the p value but also lists other useful information such as the number of genes in the category, the average log fold change, a “strength”" measure (see below) and the name with which Reactome describes the Path.
We call the function for all Paths in our incidence matrix and collect the results in a data frame:
reactomeResult <- do.call( rbind, lapply( rownames(incm), testCategory ) )As we performed many tests, we should again use a multiple testing adjustment.reactomeResult$padjust <- p.adjust( reactomeResult$pvalue, "BH" )This is a list of Reactome Paths which are significantly differentially expressed in our comparison of DPN treatment with control, sorted according to sign and strength of the signal:
reactomeResultSignif <- reactomeResult[ reactomeResult$padjust < 0.05, ]
head( reactomeResultSignif[ order(-reactomeResultSignif$strength), ] )## reactomeID numGenes avgLFC sdLFC zValue strength pvalue
## 174 191273 21 0.10808 0.09894 1.0923 0.4953 6.786e-05
## 308 381070 41 0.07055 0.13898 0.5076 0.4517 2.341e-03
## 60 1638091 30 0.06877 0.10742 0.6402 0.3767 1.499e-03
## 150 1799339 75 0.04083 0.07791 0.5240 0.3536 2.153e-05
## 405 72689 64 0.02905 0.07284 0.3989 0.2324 2.213e-03
## 23 1169091 53 -0.02579 0.06195 -0.4162 -0.1877 3.804e-03
## reactomeName padjust
## 174 Homo sapiens: Cholesterol biosynthesis 0.007838
## 308 Homo sapiens: IRE1alpha activates chaperones 0.041597
## 60 Homo sapiens: Heparan sulfate/heparin (HS-GAG) metabolism 0.032968
## 150 Homo sapiens: SRP-dependent cotranslational protein targeting to membrane 0.003316
## 405 Homo sapiens: Formation of a pool of free 40S subunits 0.041130
## 23 Homo sapiens: Activation of NF-kappaB in B cells 0.048870Working with rlog-transformed data
The rlog transform
Many common statistical methods for exploratory analysis of multidimensional data, especially methods for clustering (e.g., principal-component analysis and the like), work best for (at least approximately) homoskedastic data; this means that the variance of an observable quantity (i.e., here, the expression strength of a gene) does not depend on the mean. In RNA-Seq data, however, variance grows with the mean. For example, if one performs PCA directly on a matrix of normalized read counts, the result typically depends only on the few most strongly expressed genes because they show the largest absolute differences between samples. A simple and often used strategy to avoid this is to take the logarithm of the normalized count values plus a small pseudocount; however, now the genes with low counts tend to dominate the results because, due to the strong Poisson noise inherent to small count values, they show the strongest relative differences between samples.
As a solution, DESeq2 offers the regularized-logarithm transformation, or rlog for short. For genes with high counts, the rlog transformation differs not much from an ordinary log2 transformation. For genes with lower counts, however, the values are shrunken towards the genes’ averages across all samples. Using an empirical Bayesian prior in the form of a ridge penalty, this is done such that the rlog-transformed data are approximately homoskedastic.
Note that the rlog transformation is provided for applications other than differential testing. For differential testing we recommend the DESeq function applied to raw counts, as described earlier in this vignette, which also takes into account the dependence of the variance of counts on the mean value during the dispersion estimation step.The function rlog returns a SummarizedExperiment object which contains the rlog-transformed values in its assay slot:
rld <- rlog( dds )
assay(rld)[ 1:3, 1:3]## SRS308866 SRS308868 SRS308870
## ENSG00000000003 9.7614 9.7285 9.8653
## ENSG00000000005 -0.9813 -0.7408 -0.9447
## ENSG00000000419 8.0647 8.0770 8.1186To show the effect of the transformation, we plot the first sample against the second, first simply using the log2 function (after adding 1, to avoid taking the log of zero), and then using the rlog-transformed values.
plot( log2( 1+counts(dds, normalized=TRUE)[, 1:2] ), col="#00000020", pch=20, cex=0.3 )
plot( assay(rld)[, 1:2], col="#00000020", pch=20, cex=0.3 )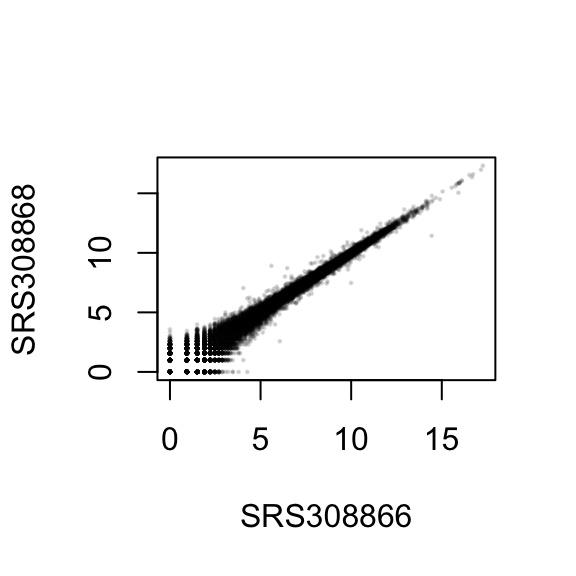
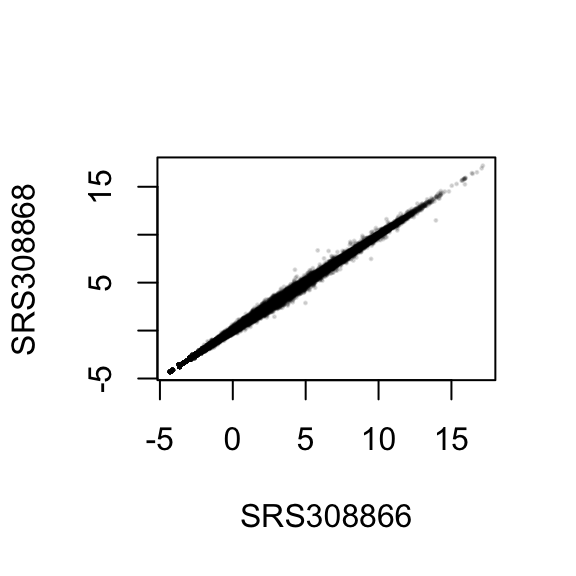
In Figure , we can see how genes with low counts seem to be excessively variable on the ordinary logarithmic scale, while the rlog transform compresses differences for genes for which the data cannot provide good information anyway.
Sample distances
A useful first step in an RNA-Seq analysis is often to assess overall similarity between samples.
We use the R function dist to calculate the Euclidean distance between samples. To avoid that the distance measure is dominated by a few highly variable genes, and have a roughly equal contribution from all genes, we use it on the rlog-transformed data:
sampleDists <- dist( t( assay(rld) ) )
as.matrix( sampleDists )[ 1:3, 1:3 ]## SRS308866 SRS308868 SRS308870
## SRS308866 0.00 32.87 31.21
## SRS308868 32.87 0.00 33.73
## SRS308870 31.21 33.73 0.00 Note the use of the function t to transpose the data matrix. We need this because dist calculates distances between data rows and our samples constitute the columns.
We visualize the distances in a heatmap, using the function heatmap.2 from the gplots package.
sampleDistMatrix <- as.matrix( sampleDists )
rownames(sampleDistMatrix) <- paste( rld$treatment,
rld$patient, sep="-" )
colnames(sampleDistMatrix) <- NULL
library( "gplots" )
library( "RColorBrewer" )
colours = colorRampPalette( rev(brewer.pal(9, "Blues")) )(255)
heatmap.2( sampleDistMatrix, trace="none", col=colours)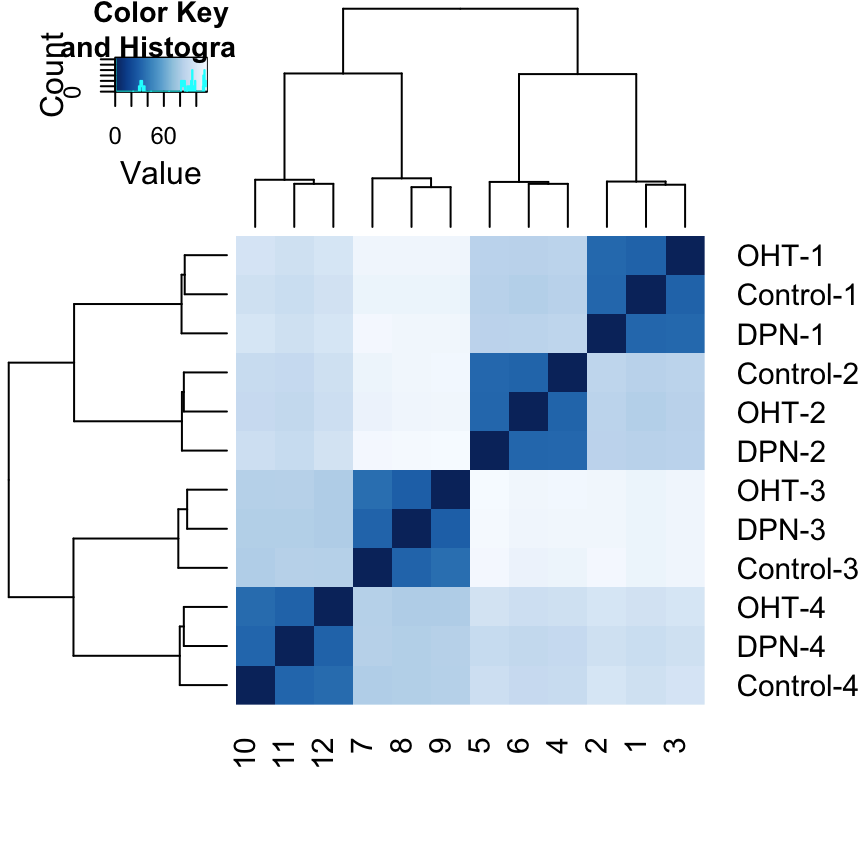
Note that we have changed the row names of the distance matrix to contain treatment type and patient number instead of sample ID, so that we have all this information in view when looking at the heatmap. Another way to visualize sample-to-sample distances is a principal-components analysis (PCA). In this ordination method, the data points (i.e., here, the samples) are projected onto the 2D plane such that they spread out optimally.
colours <- c(rgb(1:3/4,0,0),rgb(0,1:3/4,0),rgb(0,0,1:3/4),rgb(1:3/4,0,1:3/4))
plotPCA( rld, intgroup = c("patient","treatment"), col=colours )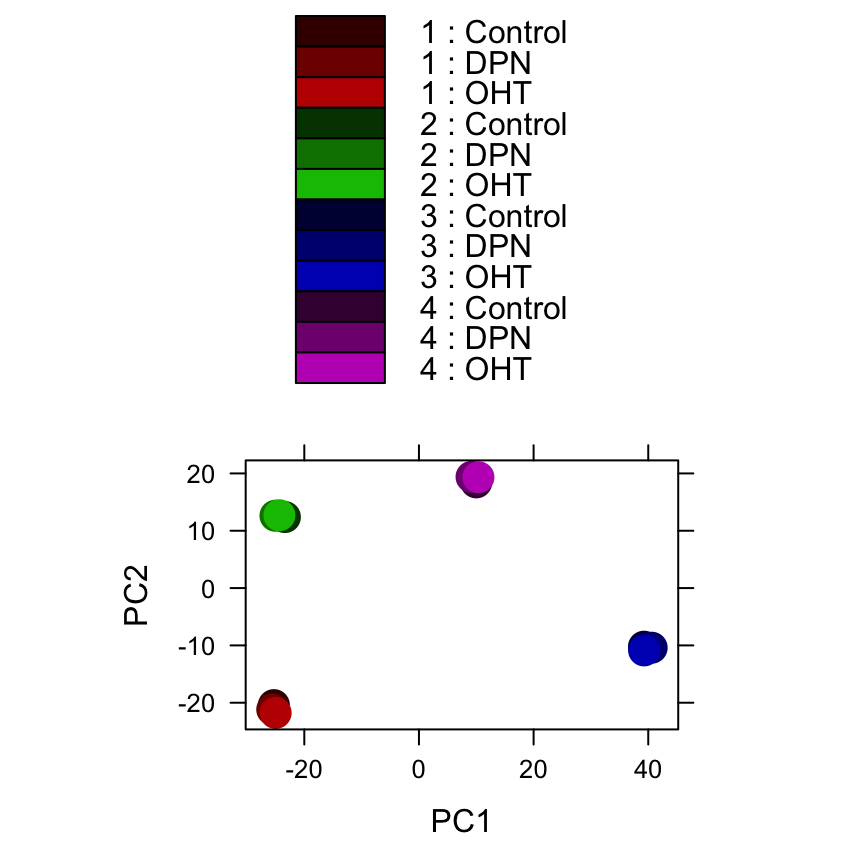
Here, we have used the function plotPCA which comes with DESeq2. The two terms specified as intgroup are column names from our sample data; they tell the function to use them to choose colours.
From both visualizations, we see that the differences between patients is much larger than the difference between treatment and control samples of the same patient. This shows why it was important to account for this paired design (``paired’’, because each treated sample is paired with one control sample from the same patient).
We did so by using the design formula ~ patient + treatment when setting up the data object in the beginning. Had we used an un-paired analysis, by specifying only , we would not have found many hits, because then, the patient-to-patient differences would have drowned out any treatment effects.
Gene clustering
In the above heatmap, the dendrogram at the side shows us a hierarchical clustering of the samples. Such a clustering can also be performed for the genes.
Since the clustering is only relevant for genes that actually carry signal, one usually carries it out only for a subset of most highly variable genes. Here, for demonstration, let us select the 35 genes with the highest variance across samples:
library( "genefilter" )
topVarGenes <- head( order( rowVars( assay(rld) ), decreasing=TRUE ), 35 )The heatmap becomes more interesting if we do not look at absolute expression strength but rather at the amount by which each gene deviates in a specific sample from the gene’s average across all samples. Hence, we center and scale each genes’ values across samples, and plot a heatmap.
heatmap.2( assay(rld)[ topVarGenes, ], scale="row",
trace="none", dendrogram="column",
col = colorRampPalette( rev(brewer.pal(9, "RdBu")) )(255),
ColSideColors = c( Control="gray", DPN="darkgreen", OHT="orange" )[
colData(rld)$treatment ] )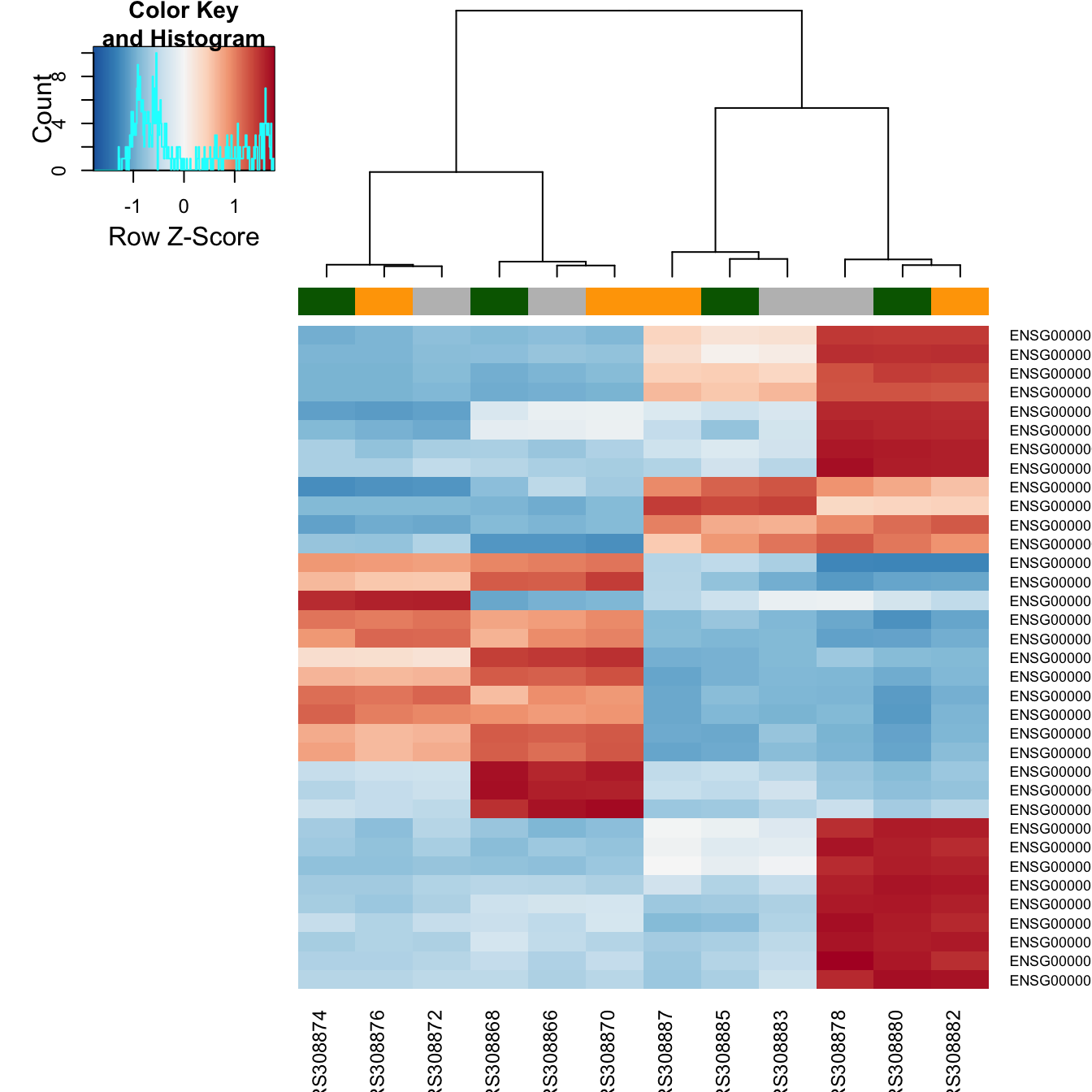
We can now see on the heatmap, blocks of genes which covary across patients. Often, such a heatmap is insightful, even though here, seeing these variations across patients is of limited value because we are rather interested in the effects between the treatments from each patient. Going further
Analyze more datasets: use the function defined in the following code chunk to download a processed count matrix from the ReCount website.
The following function takes a name of the dataset from the ReCount website, e.g. “hammer”, and returns a SummarizedExperiment object.
recount2SE <- function(name) {
filename <- paste0(name,"_eset.RData")
if (!file.exists(filename)) download.file(paste0(
"http://bowtie-bio.sourceforge.net/recount/ExpressionSets/",
filename),filename)
load(filename)
e <- get(paste0(name,".eset"))
se <- SummarizedExperiment(SimpleList(counts=exprs(e)),
colData=DataFrame(pData(e)))
se
}See also
For a more in-depth explanation of the advanced details, we advise you to proceed to the vignette of the DESeq2 package package, Differential analysis of count data. For a treatment of exon-level differential expression, we refer to the vignette of the DEXSeq package, Analyzing RN-seq data for differential exon usage with the DEXSeq package.
DEXSeq for differential exon usage. See the accompanying vignette, Analyzing RNA-seq data for differential exon usage with the DEXSeq package, which is similar to the style of this tutorial.
- Other Bioconductor packages for RNA-Seq differential expression:
edgeR, limma, DSS, BitSeq (transcript level), EBSeq, cummeRbund (for importing and visualizing Cufflinks results), monocle (single-cell analysis). More at http://bioconductor.org/packages/release/BiocViews.html#___RNASeq
- Methods for gene set testing: romer and roast in limma, permutation based: safe
- Packages for normalizing for covariates (e.g., GC content): cqn, EDASeq
- Packages for detecting batches: sva, RUVSeq
- Generating HTML results tables with links to outside resources (gene descriptions): ReportingTools
Session Info
As last part of this document, we call the function , which reports the version numbers of R and all the packages used in this session. It is good practice to always keep such a record as it will help to trace down what has happened in case that an R script ceases to work because a package has been changed in a newer version.
R version 3.1.0 (2014-04-10) Platform: x86_64-apple-darwin13.1.0 (64-bit)
locale: [1] fr_FR.UTF-8/fr_FR.UTF-8/fr_FR.UTF-8/C/fr_FR.UTF-8/fr_FR.UTF-8
attached base packages: [1] parallel stats graphics grDevices utils datasets methods base
other attached packages: [1] genefilter_1.46.1 RColorBrewer_1.0-5 gplots_2.14.2 reactome.db_1.48.0 [5] org.Hs.eg.db_2.14.0 RSQLite_0.11.4 DBI_0.3.1 DESeq2_1.4.5 [9] RcppArmadillo_0.4.450.1.0 Rcpp_0.11.3 GenomicAlignments_1.0.6 BSgenome_1.32.0 [13] GenomicFeatures_1.16.2 AnnotationDbi_1.26.0 Biobase_2.24.0 Rsamtools_1.16.1 [17] Biostrings_2.32.1 XVector_0.4.0 parathyroidSE_1.2.0 GenomicRanges_1.16.4 [21] GenomeInfoDb_1.0.2 IRanges_1.22.10 BiocGenerics_0.10.0
loaded via a namespace (and not attached): [1] annotate_1.42.1 base64enc_0.1-2 BatchJobs_1.4 BBmisc_1.7 BiocParallel_0.6.1 biomaRt_2.20.0 [7] bitops_1.0-6 brew_1.0-6 caTools_1.17.1 checkmate_1.4 codetools_0.2-9 digest_0.6.4 [13] evaluate_0.5.5 fail_1.2 foreach_1.4.2 formatR_1.0 gdata_2.13.3 geneplotter_1.42.0 [19] grid_3.1.0 gtools_3.4.1 htmltools_0.2.6 iterators_1.0.7 KernSmooth_2.23-13 knitr_1.6 [25] lattice_0.20-29 locfit_1.5-9.1 RCurl_1.95-4.3 rmarkdown_0.3.3 rtracklayer_1.24.2 sendmailR_1.2-1 [31] splines_3.1.0 stats4_3.1.0 stringr_0.6.2 survival_2.37-7 tools_3.1.0 XML_3.98-1.1 [37] xtable_1.7-4 yaml_2.1.13 zlibbioc_1.10.0
References
- Michael Love, Simon Anders, Wolfgang Huber, RNA-Seq differential expression workfow : http://www.bioconductor.org/help/course-materials/2014/BioC2014/RNA-Seq-Analysis-Lab.pdf
- Courses: http://www.bioconductor.org/help/course-materials/2014/CSAMA2014/
Show me some love with the like buttons below... Thank you and please don't forget to share and comment below!!
Montrez-moi un peu d'amour avec les like ci-dessous ... Merci et n'oubliez pas, s'il vous plaît, de partager et de commenter ci-dessous!
Recommended for You!
Recommended for you
This section contains the best data science and self-development resources to help you on your path.
Books - Data Science
Our Books
- Practical Guide to Cluster Analysis in R by A. Kassambara (Datanovia)
- Practical Guide To Principal Component Methods in R by A. Kassambara (Datanovia)
- Machine Learning Essentials: Practical Guide in R by A. Kassambara (Datanovia)
- R Graphics Essentials for Great Data Visualization by A. Kassambara (Datanovia)
- GGPlot2 Essentials for Great Data Visualization in R by A. Kassambara (Datanovia)
- Network Analysis and Visualization in R by A. Kassambara (Datanovia)
- Practical Statistics in R for Comparing Groups: Numerical Variables by A. Kassambara (Datanovia)
- Inter-Rater Reliability Essentials: Practical Guide in R by A. Kassambara (Datanovia)
Others
- R for Data Science: Import, Tidy, Transform, Visualize, and Model Data by Hadley Wickham & Garrett Grolemund
- Hands-On Machine Learning with Scikit-Learn, Keras, and TensorFlow: Concepts, Tools, and Techniques to Build Intelligent Systems by Aurelien Géron
- Practical Statistics for Data Scientists: 50 Essential Concepts by Peter Bruce & Andrew Bruce
- Hands-On Programming with R: Write Your Own Functions And Simulations by Garrett Grolemund & Hadley Wickham
- An Introduction to Statistical Learning: with Applications in R by Gareth James et al.
- Deep Learning with R by François Chollet & J.J. Allaire
- Deep Learning with Python by François Chollet
Click to follow us on Facebook :
Comment this article by clicking on "Discussion" button (top-right position of this page)







