Eigenvalues: Quick data visualization with factoextra - R software and data mining
Description
This article describes how to extract and visualize the eigenvalues/variances of the dimensions from the results of Principal Component Analysis (PCA), Correspondence Analysis (CA) and Multiple Correspondence Analysis (MCA) functions.
The R software and factoextra package are used. The functions described here are:
- get_eig() (or get_eigenvalue()): Extract the eigenvalues/variances of the principal dimensions
- fviz_eig() (or fviz_screeplot()): Plot the eigenvalues/variances against the number of dimensions
Install and load factoextra
The package devtools is required for the installation as factoextra is hosted on github.
# install.packages("devtools")
library("devtools")
install_github("kassambara/factoextra")Load factoextra :
library("factoextra")Usage
# Extraction of the eigenvalues/variances
get_eig(X)
# Visualization of the eigenvalues/variances
fviz_eig(X, choice = c("variance", "eigenvalue"),
geom = c("bar", "line"), barfill = "steelblue",
barcolor = "steelblue", linecolor = "black",
ncp = 5, addlabels = FALSE, ...)
# Alias of get_eig()
get_eigenvalue(X)
# Alias of fviz_eig()
fviz_screeplot(...)Arguments
| Argument | Description |
|---|---|
| X | an object of class PCA, CA and MCA [FactoMineR]; prcomp and princomp [stats]; dudi, pca, coa and acm [ade4]; ca and mjca [ca package]. |
| choice | a text specifying the data to be plotted. Allowed values are “variance” or “eigenvalue”. |
| geom | a text specifying the geometry to be used for the graph. Allowed values are “bar” for barplot, “line” for lineplot or c(“bar”, “line”) to use both types. |
| barfill | fill color for bar plot. |
| barcolor | outline color for bar plot. |
| linecolor | color for line plot (when geom contains “line”). |
| ncp | a numeric value specifying the number of dimensions to be shown. |
| addlabels | logical value. If TRUE, labels are added at the top of bars or points showing the information retained by each dimension. |
| … | optional arguments to be passed to the functions geom_bar(), geom_line(), geom_text() or fviz_eig(). |
Value
- get_eig() (or get_eigenvalue()): returns a data.frame containing 3 columns: the eigenvalues, the percentage of variance and the cumulative percentage of variance retained by each dimension.
- fviz_eig() (or fviz_screeplot()): returns a ggplot2 plot.
Examples
Principal Component Analysis
A Principal Component Analysis (PCA) is performed using the built-in R function prcomp() and iris data:
data(iris)
head(iris) Sepal.Length Sepal.Width Petal.Length Petal.Width Species
1 5.1 3.5 1.4 0.2 setosa
2 4.9 3.0 1.4 0.2 setosa
3 4.7 3.2 1.3 0.2 setosa
4 4.6 3.1 1.5 0.2 setosa
5 5.0 3.6 1.4 0.2 setosa
6 5.4 3.9 1.7 0.4 setosa# The variable Species (index = 5) is removed
# before the PCA analysis
res.pca <- prcomp(iris[, -5], scale = TRUE)
# Extract the eigenvalues/variances
get_eig(res.pca) eigenvalue variance.percent cumulative.variance.percent
Dim.1 2.91849782 72.9624454 72.96245
Dim.2 0.91403047 22.8507618 95.81321
Dim.3 0.14675688 3.6689219 99.48213
Dim.4 0.02071484 0.5178709 100.00000Visualize the eigenvalues/variances of the dimensions
# Default plot
fviz_eig(res.pca)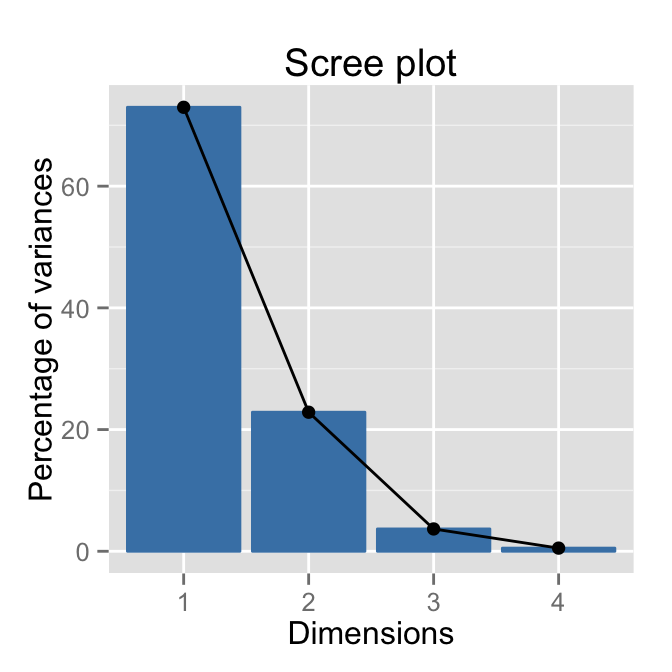
# Add labels
fviz_eig(res.pca, addlabels=TRUE, hjust = -0.3)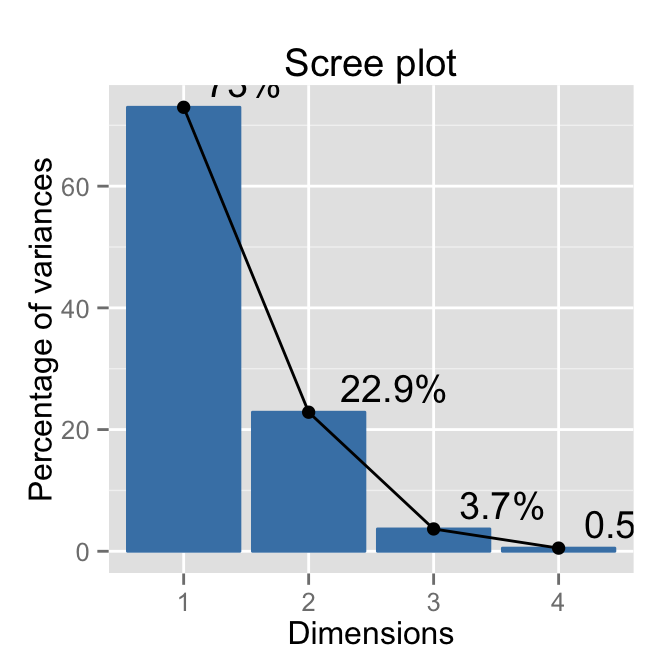
# Change the y axis limits
fviz_eig(res.pca, addlabels=TRUE, hjust = -0.3) +
ylim(0, 80)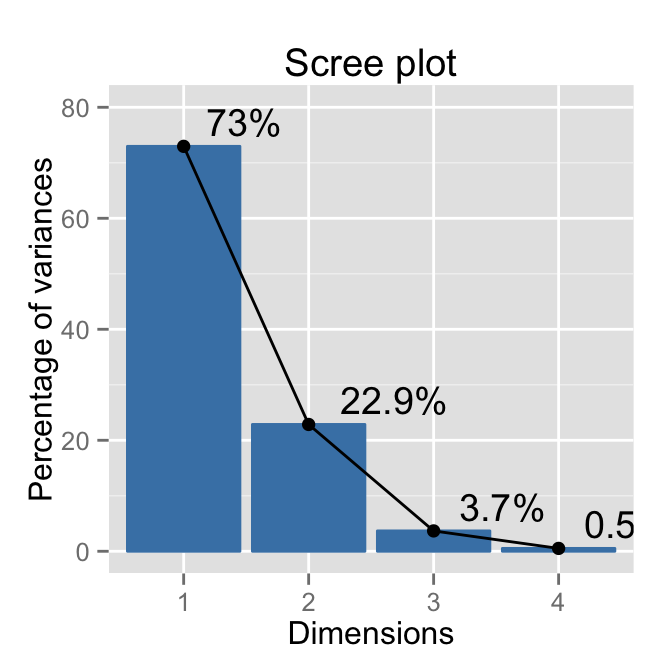
# Scree plot - Eigenvalues
fviz_eig(res.pca, choice = "eigenvalue",
addlabels=TRUE)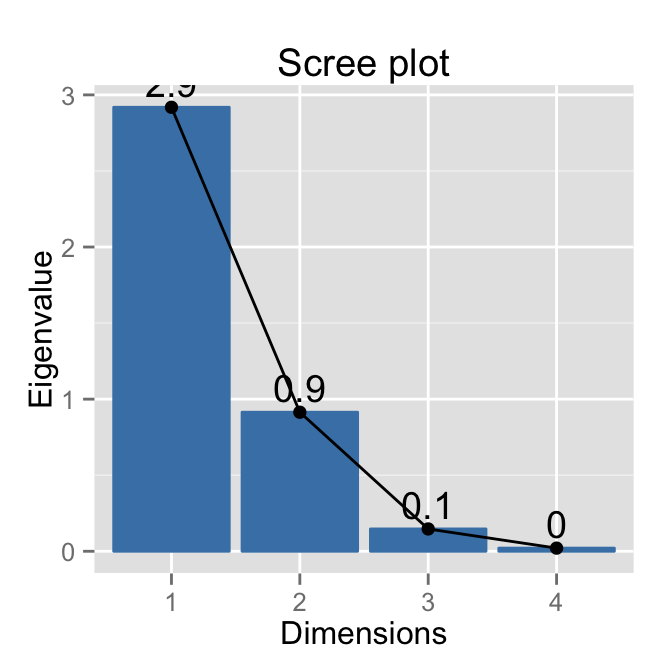
# Use only barplot
fviz_eig(res.pca, geom="bar", width=0.8, addlabels=T)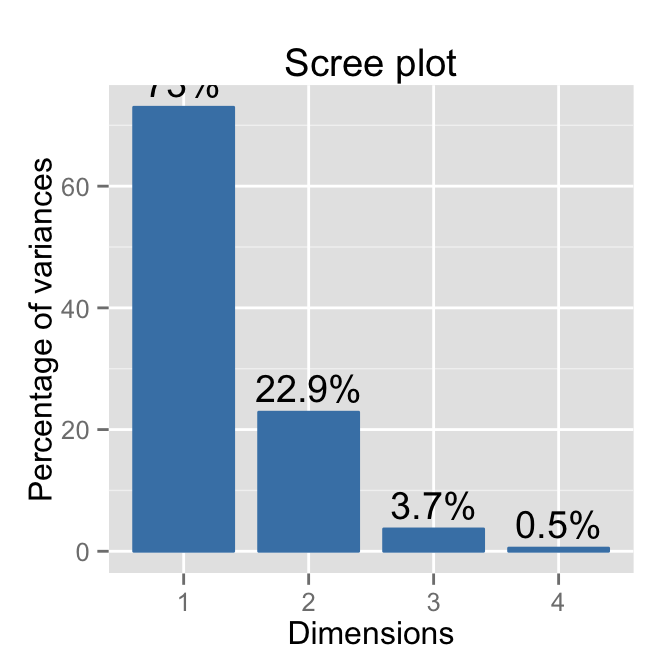
# Use only lineplot
fviz_eig(res.pca, geom="line")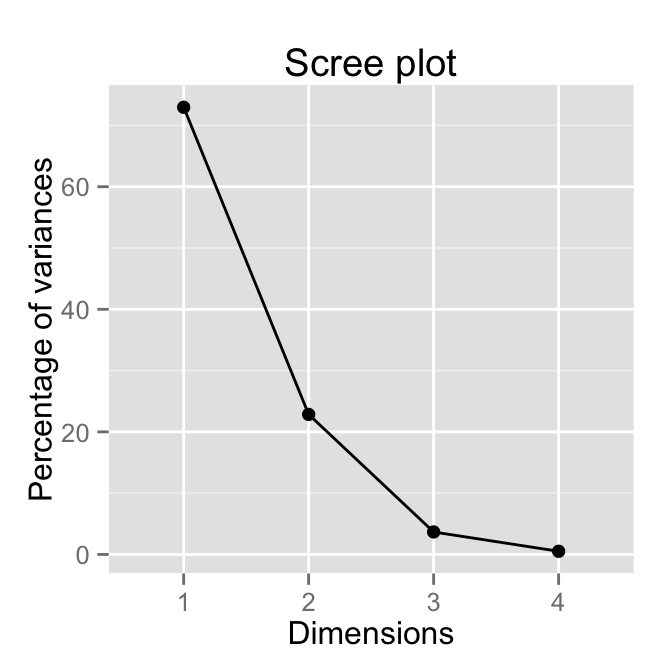
# Change theme
fviz_eig(res.pca) + theme_minimal()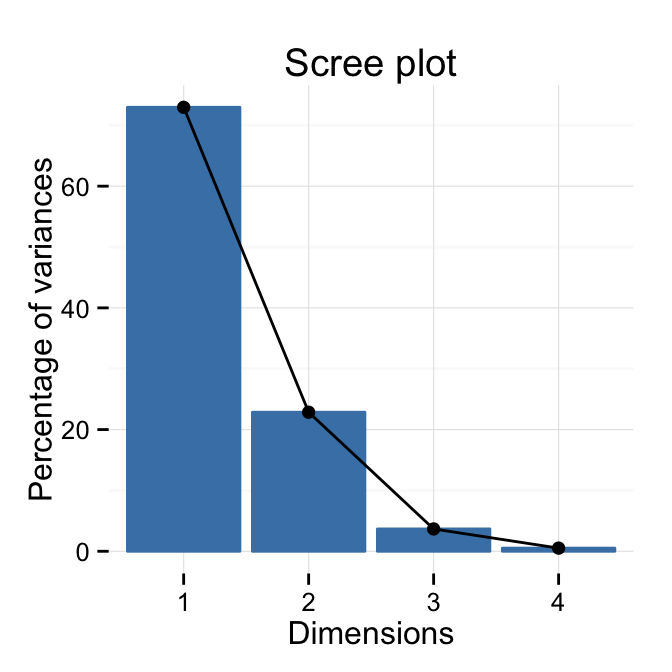
# theme_classic()
fviz_eig(res.pca) + theme_classic()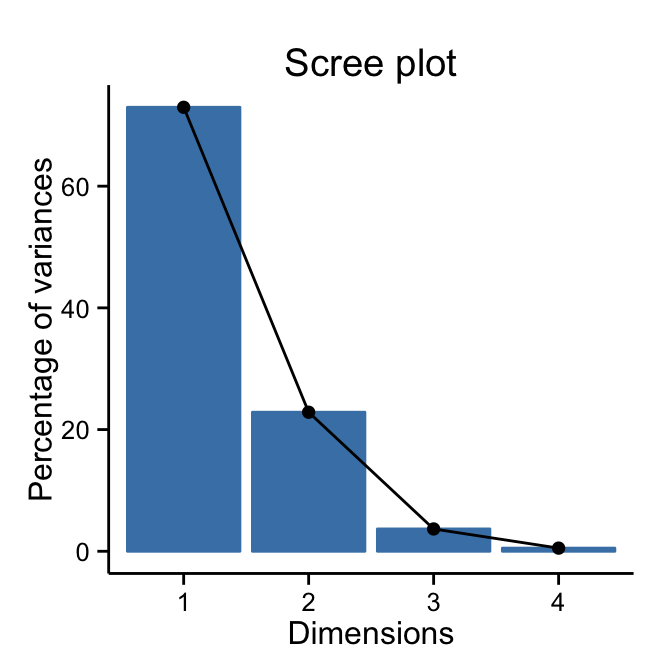
# Customized plot
fviz_eig(res.pca, addlabels=TRUE, hjust = -0.3,
linecolor ="red") + theme_minimal()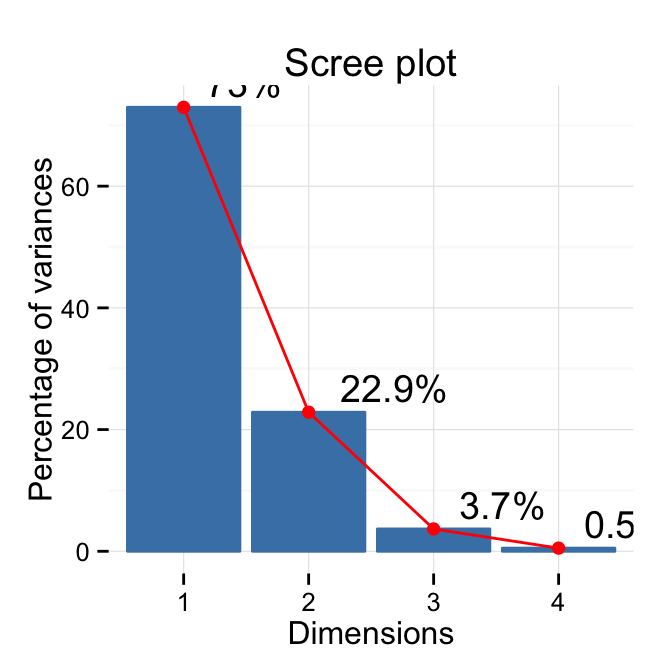
# Change colors, y axis limits and theme
p <- fviz_eig(res.pca, addlabels=TRUE, hjust = -0.3,
barfill="white", barcolor ="darkblue",
linecolor ="red") + ylim(0, 85) +
theme_minimal()
print(p)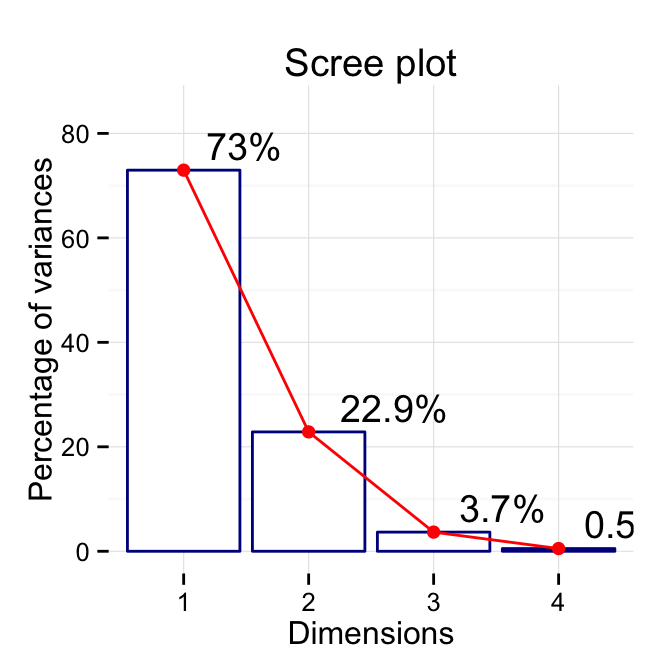
# Change titles
p + labs(title = "Variances - PCA",
x = "Principal Components", y = "% of variances")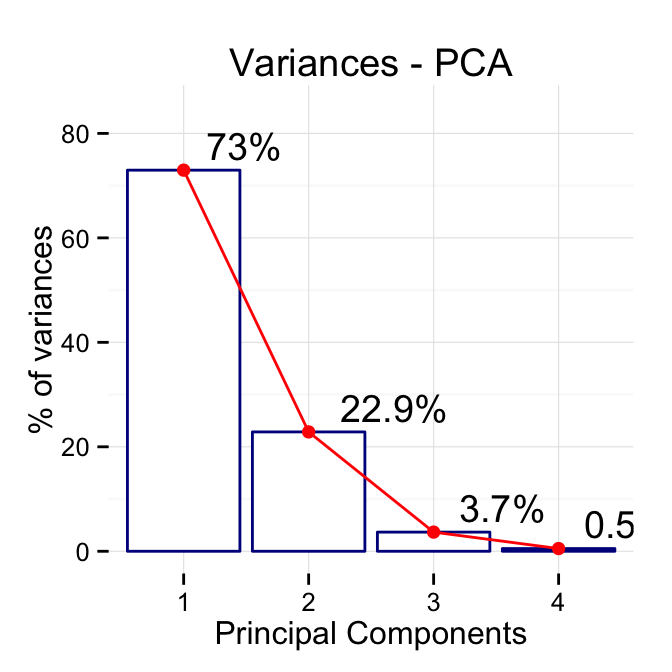
The following themes are available: theme_gray(), theme_bw(), theme_linedraw(), theme_light(), theme_minimal(), theme_classic().
Correspondence Analysis
The function CA() in FactoMineR package is used:
library(FactoMineR)
data(housetasks)
res.ca <- CA(housetasks, graph = FALSE)
get_eig(res.ca) eigenvalue variance.percent cumulative.variance.percent
Dim.1 5.428893e-01 4.869222e+01 48.69222
Dim.2 4.450028e-01 3.991269e+01 88.60491
Dim.3 1.270484e-01 1.139509e+01 100.00000
Dim.4 5.119700e-33 4.591904e-31 100.00000fviz_eig(res.ca)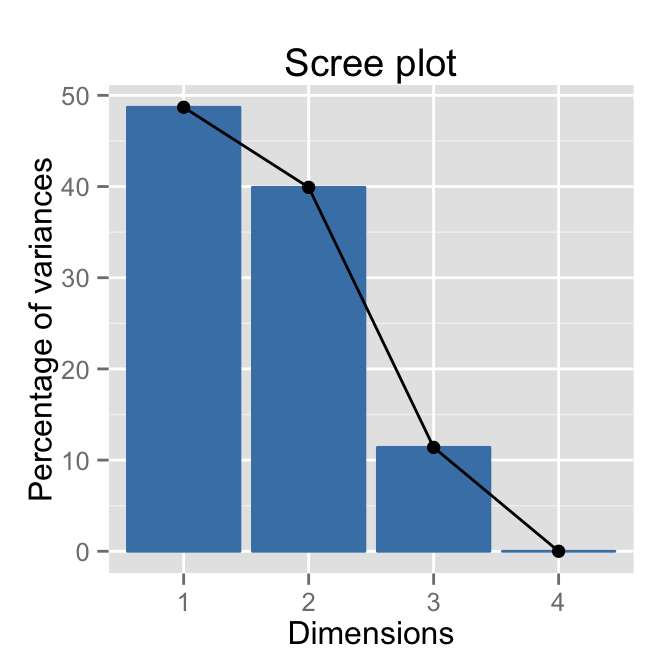
Multiple Correspondence Analysis
The function MCA() in FactoMineR package is used:
library(FactoMineR)
data(poison)
res.mca <- MCA(poison, quanti.sup = 1:2,
quali.sup = 3:4, graph=FALSE)
get_eig(res.mca) eigenvalue variance.percent cumulative.variance.percent
Dim.1 0.33523140 33.523140 33.52314
Dim.2 0.12913979 12.913979 46.43712
Dim.3 0.10734849 10.734849 57.17197
Dim.4 0.09587950 9.587950 66.75992
Dim.5 0.07883277 7.883277 74.64319
Dim.6 0.07108981 7.108981 81.75217
Dim.7 0.06016580 6.016580 87.76876
Dim.8 0.05577301 5.577301 93.34606
Dim.9 0.04120578 4.120578 97.46663
Dim.10 0.01304158 1.304158 98.77079
Dim.11 0.01229208 1.229208 100.00000fviz_eig(res.mca, ncp = 10)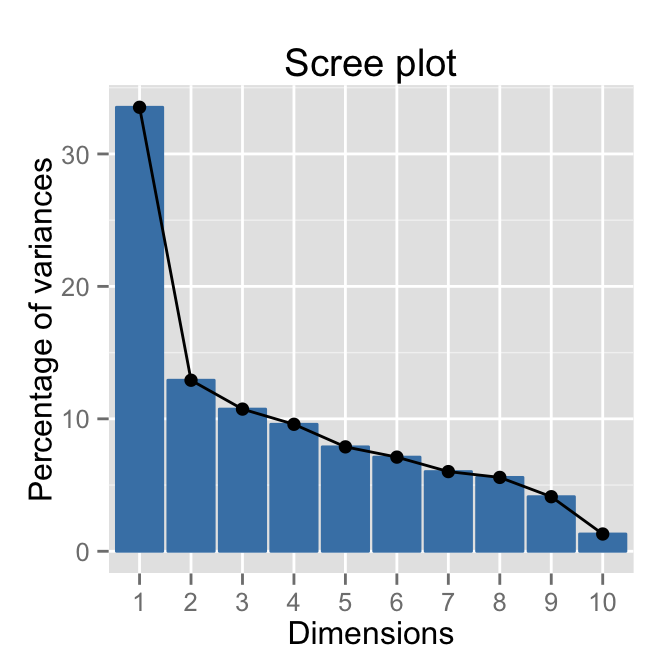
Infos
This analysis has been performed using R software (ver. 3.1.2) and factoextra (ver. 1.0.2)
Show me some love with the like buttons below... Thank you and please don't forget to share and comment below!!
Montrez-moi un peu d'amour avec les like ci-dessous ... Merci et n'oubliez pas, s'il vous plaît, de partager et de commenter ci-dessous!
Recommended for You!
Recommended for you
This section contains the best data science and self-development resources to help you on your path.
Books - Data Science
Our Books
- Practical Guide to Cluster Analysis in R by A. Kassambara (Datanovia)
- Practical Guide To Principal Component Methods in R by A. Kassambara (Datanovia)
- Machine Learning Essentials: Practical Guide in R by A. Kassambara (Datanovia)
- R Graphics Essentials for Great Data Visualization by A. Kassambara (Datanovia)
- GGPlot2 Essentials for Great Data Visualization in R by A. Kassambara (Datanovia)
- Network Analysis and Visualization in R by A. Kassambara (Datanovia)
- Practical Statistics in R for Comparing Groups: Numerical Variables by A. Kassambara (Datanovia)
- Inter-Rater Reliability Essentials: Practical Guide in R by A. Kassambara (Datanovia)
Others
- R for Data Science: Import, Tidy, Transform, Visualize, and Model Data by Hadley Wickham & Garrett Grolemund
- Hands-On Machine Learning with Scikit-Learn, Keras, and TensorFlow: Concepts, Tools, and Techniques to Build Intelligent Systems by Aurelien Géron
- Practical Statistics for Data Scientists: 50 Essential Concepts by Peter Bruce & Andrew Bruce
- Hands-On Programming with R: Write Your Own Functions And Simulations by Garrett Grolemund & Hadley Wickham
- An Introduction to Statistical Learning: with Applications in R by Gareth James et al.
- Deep Learning with R by François Chollet & J.J. Allaire
- Deep Learning with Python by François Chollet
Click to follow us on Facebook :
Comment this article by clicking on "Discussion" button (top-right position of this page)







