Visualize Multiple Factor Analysis
fviz_mfa.RdMultiple factor analysis (MFA) is used to analyze a data set in
which individuals are described by several sets of variables (quantitative
and/or qualitative) structured into groups. fviz_mfa() provides
ggplot2-based elegant visualization of MFA outputs from the R function: MFA
[FactoMineR].
fviz_mfa_ind(): Graph of individuals
fviz_mfa_var(): Graph of variables
fviz_mfa_axes(): Graph of partial axes
fviz_mfa(): An alias of fviz_mfa_ind(res.mfa, partial = "all")
fviz_mfa_quali_biplot(): Biplot of individuals and qualitative variables
fviz_mfa_ind(X, axes = c(1, 2), geom = c("point", "text"), repel = FALSE, habillage = "none", palette = NULL, addEllipses = FALSE, col.ind = "blue", col.ind.sup = "darkblue", alpha.ind = 1, shape.ind = 19, col.quali.var.sup = "black", select.ind = list(name = NULL, cos2 = NULL, contrib = NULL), partial = NULL, col.partial = "group", ...) fviz_mfa_quali_biplot(X, axes = c(1, 2), geom = c("point", "text"), repel = repel, title = "Biplot of individuals and qualitative variables - MFA", ...) fviz_mfa_var(X, choice = c("quanti.var", "group", "quali.var"), axes = c(1, 2), geom = c("point", "text"), repel = FALSE, habillage = "none", col.var = "red", alpha.var = 1, shape.var = 17, col.var.sup = "darkgreen", palette = NULL, select.var = list(name = NULL, cos2 = NULL, contrib = NULL), ...) fviz_mfa_axes(X, axes = c(1, 2), geom = c("arrow", "text"), col.axes = NULL, alpha.axes = 1, col.circle = "grey70", select.axes = list(name = NULL, contrib = NULL), repel = FALSE, ...) fviz_mfa(X, partial = "all", ...)
Arguments
| X | an object of class MFA [FactoMineR]. |
|---|---|
| axes | a numeric vector of length 2 specifying the dimensions to be plotted. |
| geom | a text specifying the geometry to be used for the graph. Allowed
values are the combination of |
| repel | a boolean, whether to use ggrepel to avoid overplotting text labels or not. |
| habillage | an optional factor variable for coloring the observations by groups. Default value is "none". If X is an MFA object from FactoMineR package, habillage can also specify the index of the factor variable in the data. |
| palette | the color palette to be used for coloring or filling by groups. Allowed values include "grey" for grey color palettes; brewer palettes e.g. "RdBu", "Blues", ...; or custom color palette e.g. c("blue", "red"); and scientific journal palettes from ggsci R package, e.g.: "npg", "aaas", "lancet", "jco", "ucscgb", "uchicago", "simpsons" and "rickandmorty". Can be also a numeric vector of length(groups); in this case a basic color palette is created using the function palette. |
| addEllipses | logical value. If TRUE, draws ellipses around the individuals when habillage != "none". |
| col.ind, col.var, col.axes | color for individuals, variables and col.axes respectively. Can be a continuous variable or a factor variable. Possible values include also : "cos2", "contrib", "coord", "x" or "y". In this case, the colors for individuals/variables are automatically controlled by their qualities ("cos2"), contributions ("contrib"), coordinates (x^2 + y^2 , "coord"), x values("x") or y values("y"). To use automatic coloring (by cos2, contrib, ....), make sure that habillage ="none". |
| col.ind.sup | color for supplementary individuals |
| alpha.ind, alpha.var, alpha.axes | controls the transparency of individual, variable, group and axes colors, respectively. The value can variate from 0 (total transparency) to 1 (no transparency). Default value is 1. Possible values include also : "cos2", "contrib", "coord", "x" or "y". In this case, the transparency for individual/variable colors are automatically controlled by their qualities ("cos2"), contributions ("contrib"), coordinates (x^2 + y^2 , "coord"), x values("x") or y values("y"). To use this, make sure that habillage ="none". |
| shape.ind, shape.var | point shapes of individuals, variables, groups and axes |
| col.quali.var.sup | color for supplementary qualitative variables. Default is "black". |
| select.ind, select.var, select.axes | a selection of individuals/partial individuals/ variables/groups/axes to be drawn. Allowed values are NULL or a list containing the arguments name, cos2 or contrib:
|
| partial | list of the individuals for which the partial points should be drawn. (by default, partial = NULL and no partial points are drawn). Use partial = "All" to visualize partial points for all individuals. |
| col.partial | color for partial individuals. By default, points are colored according to the groups. |
| ... | Arguments to be passed to the function fviz() |
| title | the title of the graph |
| choice | the graph to plot. Allowed values include one of c("quanti.var", "quali.var", "group") for plotting quantitative variables, qualitative variables and group of variables, respectively. |
| col.var.sup | color for supplementary variables. |
| col.circle | a color for the correlation circle. Used only when X is a PCA output. |
Value
a ggplot2 plot
References
http://www.sthda.com/english/
Examples
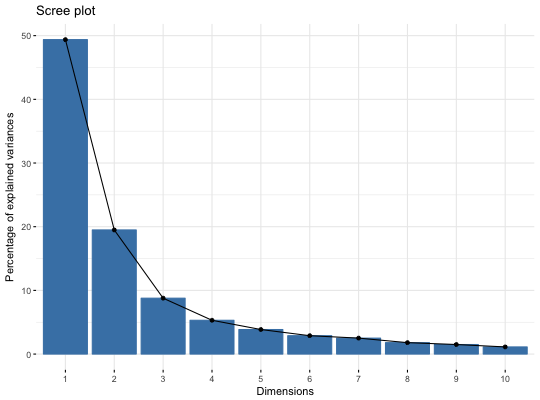
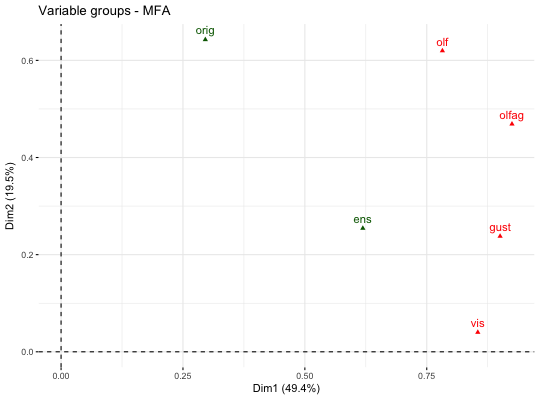
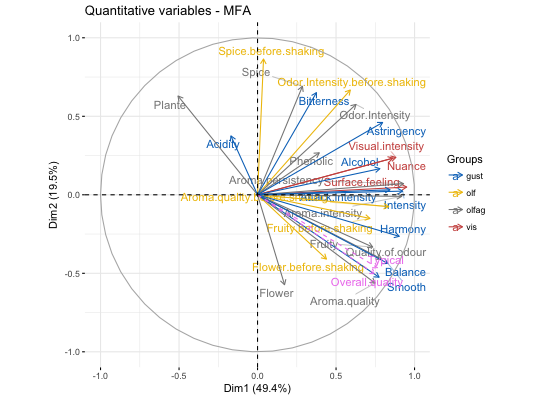
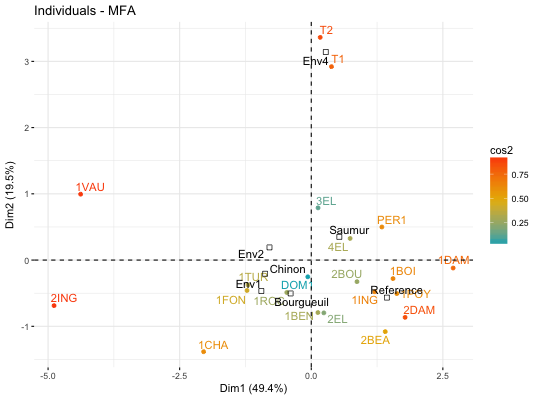
# Compute Multiple Factor Analysis library("FactoMineR") data(wine) res.mfa <- MFA(wine, group=c(2,5,3,10,9,2), type=c("n",rep("s",5)), ncp=5, name.group=c("orig","olf","vis","olfag","gust","ens"), num.group.sup=c(1,6), graph=FALSE) # Eigenvalues/variances of dimensions fviz_screeplot(res.mfa)# Group of variables fviz_mfa_var(res.mfa, "group")# Quantitative variables fviz_mfa_var(res.mfa, "quanti.var", palette = "jco", col.var.sup = "violet", repel = TRUE)# Graph of individuals colored by cos2 fviz_mfa_ind(res.mfa, col.ind = "cos2", gradient.cols = c("#00AFBB", "#E7B800", "#FC4E07"), repel = TRUE)# Partial individuals fviz_mfa_ind(res.mfa, partial = "all")# Partial axes fviz_mfa_axes(res.mfa)# Graph of categorical variable categories # ++++++++++++++++++++++++++++++++++++++++ data(poison) res.mfa <- MFA(poison, group=c(2,2,5,6), type=c("s","n","n","n"), name.group=c("desc","desc2","symptom","eat"), num.group.sup=1:2, graph=FALSE) # Plot of qualitative variables fviz_mfa_var(res.mfa, "quali.var")# Biplot of categorical variable categories and individuals # +++++++++++++++++++++++++++++++++++++++++++++++++++++++++ # Use repel = TRUE to avoid overplotting grp <- as.factor(poison[, "Vomiting"]) fviz_mfa_quali_biplot(res.mfa, repel = FALSE, col.var = "#E7B800", habillage = grp, addEllipses = TRUE, ellipse.level = 0.95)