In this tutorial, you will learn how to select or subset data frame columns by names and position using the R function select() and pull() [in dplyr package]. We’ll also show how to remove columns from a data frame.
You will learn how to use the following functions:
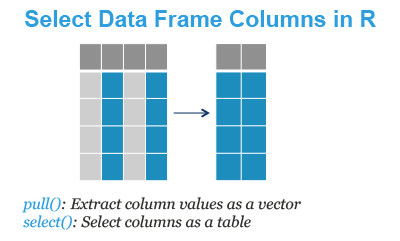
- pull(): Extract column values as a vector. The column of interest can be specified either by name or by index.
- select(): Extract one or multiple columns as a data table. It can be also used to remove columns from the data frame.
- select_if(): Select columns based on a particular condition. One can use this function to, for example, select columns if they are numeric.
- Helper functions - starts_with(), ends_with(), contains(), matches(), one_of(): Select columns/variables based on their names
Contents:
Required packages
Load the tidyverse packages, which include dplyr:
library(tidyverse)Demo dataset
We’ll use the R built-in iris data set, which we start by converting into a tibble data frame (tbl_df) for easier data analysis.
my_data <- as_tibble(iris)
my_data## # A tibble: 150 x 5
## Sepal.Length Sepal.Width Petal.Length Petal.Width Species
## <dbl> <dbl> <dbl> <dbl> <fct>
## 1 5.1 3.5 1.4 0.2 setosa
## 2 4.9 3 1.4 0.2 setosa
## 3 4.7 3.2 1.3 0.2 setosa
## 4 4.6 3.1 1.5 0.2 setosa
## 5 5 3.6 1.4 0.2 setosa
## 6 5.4 3.9 1.7 0.4 setosa
## # ... with 144 more rowsExtract column values as a vector
my_data %>% pull(Species)## [1] setosa setosa setosa setosa setosa setosa
## [7] setosa setosa setosa setosa setosa setosa
## [13] setosa setosa setosa setosa setosa setosa
## [19] setosa setosa setosa setosa setosa setosa
## [25] setosa setosa setosa setosa setosa setosa
## [31] setosa setosa setosa setosa setosa setosa
## [37] setosa setosa setosa setosa setosa setosa
## [43] setosa setosa setosa setosa setosa setosa
## [49] setosa setosa versicolor versicolor versicolor versicolor
## [55] versicolor versicolor versicolor versicolor versicolor versicolor
## [61] versicolor versicolor versicolor versicolor versicolor versicolor
## [67] versicolor versicolor versicolor versicolor versicolor versicolor
## [73] versicolor versicolor versicolor versicolor versicolor versicolor
## [79] versicolor versicolor versicolor versicolor versicolor versicolor
## [85] versicolor versicolor versicolor versicolor versicolor versicolor
## [91] versicolor versicolor versicolor versicolor versicolor versicolor
## [97] versicolor versicolor versicolor versicolor virginica virginica
## [103] virginica virginica virginica virginica virginica virginica
## [109] virginica virginica virginica virginica virginica virginica
## [115] virginica virginica virginica virginica virginica virginica
## [121] virginica virginica virginica virginica virginica virginica
## [127] virginica virginica virginica virginica virginica virginica
## [133] virginica virginica virginica virginica virginica virginica
## [139] virginica virginica virginica virginica virginica virginica
## [145] virginica virginica virginica virginica virginica virginica
## Levels: setosa versicolor virginicaExtract columns as a data table
Select column by position
- Select columns 1 to 3:
my_data %>% select(1:3)- Select column 1 and 3 but not 2:
my_data %>% select(1, 3)Select columns by names
Select columns by names: Sepal.Length and Petal.Length
my_data %>% select(Sepal.Length, Petal.Length)## # A tibble: 150 x 2
## Sepal.Length Petal.Length
## <dbl> <dbl>
## 1 5.1 1.4
## 2 4.9 1.4
## 3 4.7 1.3
## 4 4.6 1.5
## 5 5 1.4
## 6 5.4 1.7
## # ... with 144 more rowsSelect all columns from Sepal.Length to Petal.Length
my_data %>% select(Sepal.Length:Petal.Length)## # A tibble: 150 x 3
## Sepal.Length Sepal.Width Petal.Length
## <dbl> <dbl> <dbl>
## 1 5.1 3.5 1.4
## 2 4.9 3 1.4
## 3 4.7 3.2 1.3
## 4 4.6 3.1 1.5
## 5 5 3.6 1.4
## 6 5.4 3.9 1.7
## # ... with 144 more rowsThere are several special functions that can be used inside select(): starts_with(), ends_with(), contains(), matches(), one_of(), etc.
# Select column whose name starts with "Petal"
my_data %>% select(starts_with("Petal"))
# Select column whose name ends with "Width"
my_data %>% select(ends_with("Width"))
# Select columns whose names contains "etal"
my_data %>% select(contains("etal"))
# Select columns whose name maches a regular expression
my_data %>% select(matches(".t."))
# selects variables provided in a character vector.
my_data %>% select(one_of(c("Sepal.Length", "Petal.Length")))Select column based on a condtion
It’s possible to apply a function to the columns. The columns for which the function returns TRUE are selected.
Select only numeric columns:
my_data %>% select_if(is.numeric)## # A tibble: 150 x 4
## Sepal.Length Sepal.Width Petal.Length Petal.Width
## <dbl> <dbl> <dbl> <dbl>
## 1 5.1 3.5 1.4 0.2
## 2 4.9 3 1.4 0.2
## 3 4.7 3.2 1.3 0.2
## 4 4.6 3.1 1.5 0.2
## 5 5 3.6 1.4 0.2
## 6 5.4 3.9 1.7 0.4
## # ... with 144 more rowsRemove columns
Note that, to remove a column from a data frame, prepend its name by minus -.
Removing Sepal.Length and Petal.Length columns:
my_data %>% select(-Sepal.Length, -Petal.Length)Removing all columns from Sepal.Length to Petal.Length:
my_data %>% select(-(Sepal.Length:Petal.Length))## # A tibble: 150 x 2
## Petal.Width Species
## <dbl> <fct>
## 1 0.2 setosa
## 2 0.2 setosa
## 3 0.2 setosa
## 4 0.2 setosa
## 5 0.2 setosa
## 6 0.4 setosa
## # ... with 144 more rowsRemoving all columns whose name starts with “Petal”:
my_data %>% select(-starts_with("Petal"))## # A tibble: 150 x 3
## Sepal.Length Sepal.Width Species
## <dbl> <dbl> <fct>
## 1 5.1 3.5 setosa
## 2 4.9 3 setosa
## 3 4.7 3.2 setosa
## 4 4.6 3.1 setosa
## 5 5 3.6 setosa
## 6 5.4 3.9 setosa
## # ... with 144 more rowsNote that, if you want to drop columns by position, the syntax is as follow.
# Drop column 1
my_data %>% select(-1)
# Drop columns 1 to 3
my_data %>% select(-(1:3))
# Drop columns 1 and 3 but not 2
my_data %>% select(-1, -3)Summary
In this tutorial, we describe how to select columns by positions and by names. Additionally, we present how to remove columns from a data frame.
Recommended for you
This section contains best data science and self-development resources to help you on your path.
Coursera - Online Courses and Specialization
Data science
- Course: Machine Learning: Master the Fundamentals by Stanford
- Specialization: Data Science by Johns Hopkins University
- Specialization: Python for Everybody by University of Michigan
- Courses: Build Skills for a Top Job in any Industry by Coursera
- Specialization: Master Machine Learning Fundamentals by University of Washington
- Specialization: Statistics with R by Duke University
- Specialization: Software Development in R by Johns Hopkins University
- Specialization: Genomic Data Science by Johns Hopkins University
Popular Courses Launched in 2020
- Google IT Automation with Python by Google
- AI for Medicine by deeplearning.ai
- Epidemiology in Public Health Practice by Johns Hopkins University
- AWS Fundamentals by Amazon Web Services
Trending Courses
- The Science of Well-Being by Yale University
- Google IT Support Professional by Google
- Python for Everybody by University of Michigan
- IBM Data Science Professional Certificate by IBM
- Business Foundations by University of Pennsylvania
- Introduction to Psychology by Yale University
- Excel Skills for Business by Macquarie University
- Psychological First Aid by Johns Hopkins University
- Graphic Design by Cal Arts
Amazon FBA
Amazing Selling Machine
Books - Data Science
Our Books
- Practical Guide to Cluster Analysis in R by A. Kassambara (Datanovia)
- Practical Guide To Principal Component Methods in R by A. Kassambara (Datanovia)
- Machine Learning Essentials: Practical Guide in R by A. Kassambara (Datanovia)
- R Graphics Essentials for Great Data Visualization by A. Kassambara (Datanovia)
- GGPlot2 Essentials for Great Data Visualization in R by A. Kassambara (Datanovia)
- Network Analysis and Visualization in R by A. Kassambara (Datanovia)
- Practical Statistics in R for Comparing Groups: Numerical Variables by A. Kassambara (Datanovia)
- Inter-Rater Reliability Essentials: Practical Guide in R by A. Kassambara (Datanovia)
Others
- R for Data Science: Import, Tidy, Transform, Visualize, and Model Data by Hadley Wickham & Garrett Grolemund
- Hands-On Machine Learning with Scikit-Learn, Keras, and TensorFlow: Concepts, Tools, and Techniques to Build Intelligent Systems by Aurelien Géron
- Practical Statistics for Data Scientists: 50 Essential Concepts by Peter Bruce & Andrew Bruce
- Hands-On Programming with R: Write Your Own Functions And Simulations by Garrett Grolemund & Hadley Wickham
- An Introduction to Statistical Learning: with Applications in R by Gareth James et al.
- Deep Learning with R by François Chollet & J.J. Allaire
- Deep Learning with Python by François Chollet



Thank you for sharing!
Your feedback is appreciated!
Thanks for the example!
One of the things I am always trying and failing to do is to work with a set of variables from a data frame – not pulling them out, but applying a function to them.
If I use the ‘select’ command, then I’ve created a copy of the data frame, and have to match it back to the original.
Is there any way to use pipes and dplyr to apply a command to a subset of columns?
Thank you for your comments.
What do you think about the mutant() family of functions described at: https://www.datanovia.com/en/lessons/compute-and-add-new-variables-to-a-data-frame-in-r/ ?
thank you for your sharing
i am beginner in R. i have some questions.
how combine multi table ( i used .csv) to be one, because i saw that from those tables (8 tables) the field name is all same, but the file name is different.
can you help me how to combine those tables and add 1 field from each table (field name is Value) and added field name change to the table name?
e.g : number of table : 8 (T1 – T8)
number of field in every table = 10 (F1 – F10). F1 – F9 in T1 – T10 fill with the same data value
Data value of F10 is the only field that filled different value in every tables
How to combine (T9 ) with F1-F9, and add F10-F17 named as the tables name and will fills with the value of F10 in every tables?
I hope you can help and give your response to me. Thank you so much
very helpful. Thanks a lot!!!
Hello, I tried the pull method on a column in my dataframe but received the following error message:
Error in UseMethod(“pull”) :
no applicable method for ‘pull’ applied to an object of class “factor”
Is there anything I can do to fix this?
Did this work?
iris %>% dplyr::pull(Species)
Thanks !!!