Description
Although there are several good books on unsupervised machine learning, we felt that many of them are too theoretical. This book provides practical guide to cluster analysis, elegant visualization and interpretation. It contains 5 parts. Part I provides a quick introduction to R and presents required R packages, as well as, data formats and dissimilarity measures for cluster analysis and visualization. Part II covers partitioning clustering methods, which subdivide the data sets into a set of k groups, where k is the number of groups pre-specified by the analyst. Partitioning clustering approaches include: K-means, K-Medoids (PAM) and CLARA algorithms. In Part III, we consider hierarchical clustering method, which is an alternative approach to partitioning clustering. The result of hierarchical clustering is a tree-based representation of the objects called dendrogram. In this part, we describe how to compute, visualize, interpret and compare dendrograms. Part IV describes clustering validation and evaluation strategies, which consists of measuring the goodness of clustering results. Among the chapters covered here, there are: Assessing clustering tendency, Determining the optimal number of clusters, Cluster validation statistics, Choosing the best clustering algorithms and Computing p-value for hierarchical clustering. Part V presents advanced clustering methods, including: Hierarchical k-means clustering, Fuzzy clustering, Model-based clustering and Density-based clustering.
Introduction
Large amounts of data are collected every day from satellite images, bio-medical, security, marketing, web search, geo-spatial or other automatic equipment. Mining knowledge from these big data far exceeds human’s abilities.
Clustering is one of the important data mining methods for discovering knowledge in multidimensional data. The goal of clustering is to identify pattern or groups of similar objects within a data set of interest.
In the litterature, it is referred as “pattern recognition” or “unsupervised machine learning” - “unsupervised” because we are not guided by a priori ideas of which variables or samples belong in which clusters. “Learning” because the machine algorithm “learns” how to cluster.
Cluster analysis is popular in many fields, including:
- In cancer research for classifying patients into subgroups according their gene expression profile. This can be useful for identifying the molecular profile of patients with good or bad prognostic, as well as for understanding the disease.
- In marketing for market segmentation by identifying subgroups of customers with similar profiles and who might be receptive to a particular form of advertising.
- In City-planning for identifying groups of houses according to their type, value and location.
Key features of this book
Although there are several good books on unsupervised machine learning/clustering and related topics, we felt that many of them are either too high-level, theoretical or too advanced. Our goal was to write a practical guide to cluster analysis, elegant visualization and interpretation.
The main parts of the book include:
- distance measures,
- partitioning clustering,
- hierarchical clustering,
- cluster validation methods, as well as,
- advanced clustering methods such as fuzzy clustering, density-based clustering and model-based clustering.
The book presents the basic principles of these tasks and provide many examples in R. This book offers solid guidance in data mining for students and researchers.
Key features:
- Covers clustering algorithm and implementation
- Key mathematical concepts are presented
- Short, self-contained chapters with practical examples. This means that, you don’t need to read the different chapters in sequence.
How this book is organized?
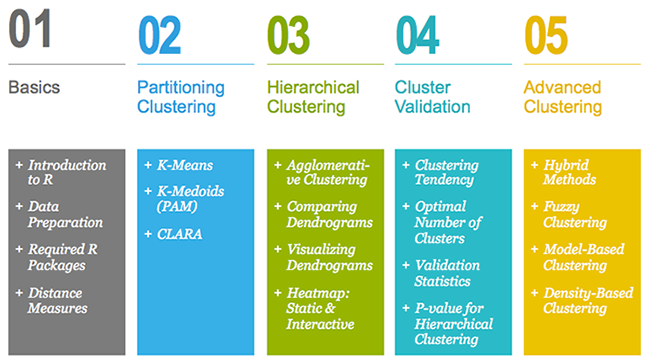
This book contains 5 parts. Part I (Chapter 1 - 3) provides a quick introduction to R (chapter 1) and presents required R packages and data format (Chapter 2) for clustering analysis and visualization.
The classification of objects, into clusters, requires some methods for measuring the distance or the (dis)similarity between the objects. Chapter 3 covers the common distance measures used for assessing similarity between observations.
Part II starts with partitioning clustering methods, which include:
- K-means clustering (Chapter 4),
- K-Medoids or PAM (partitioning around medoids) algorithm (Chapter 5) and
- CLARA algorithms (Chapter 6).
Partitioning clustering approaches subdivide the data sets into a set of k groups, where k is the number of groups pre-specified by the analyst.
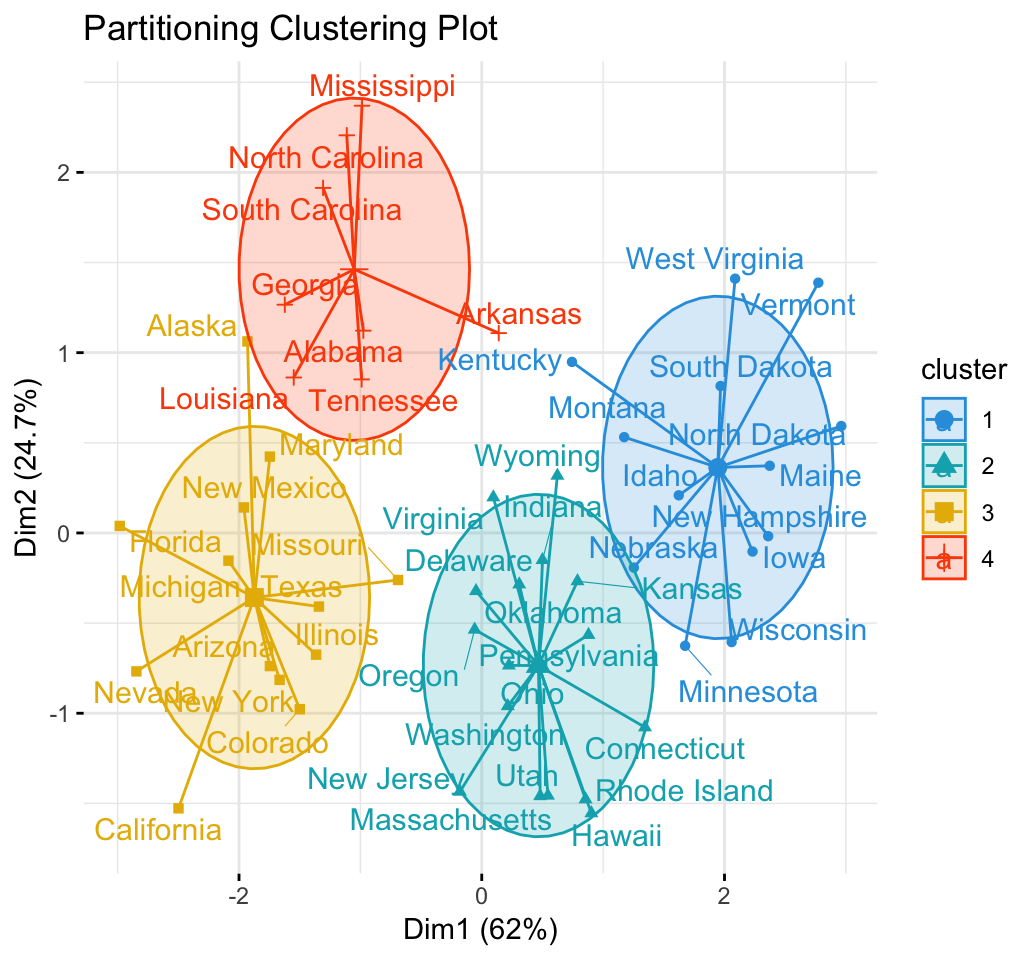
In Part III, we consider agglomerative hierarchical clustering method, which is an alternative approach to partitionning clustering for identifying groups in a data set. It does not require to pre-specify the number of clusters to be generated. The result of hierarchical clustering is a tree-based representation of the objects, which is also known as dendrogram (see the figure below).
In this part, we describe how to compute, visualize, interpret and compare dendrograms:
- Agglomerative clustering (Chapter 7)
- Algorithm and steps
- Verify the cluster tree
- Cut the dendrogram into different groups
- Compare dendrograms (Chapter 8)
- Visual comparison of two dendrograms
- Correlation matrix between a list of dendrograms
- Visualize dendrograms (Chapter 9)
- Case of small data sets
- Case of dendrogram with large data sets: zoom, sub-tree, PDF
- Customize dendrograms using dendextend
- Heatmap: static and interactive (Chapter 10)
- R base heat maps
- Pretty heat maps
- Interactive heat maps
- Complex heatmap
- Real application: gene expression data
In this section, you will learn how to generate and interpret the following plots.
- Standard dendrogram with filled rectangle around clusters:
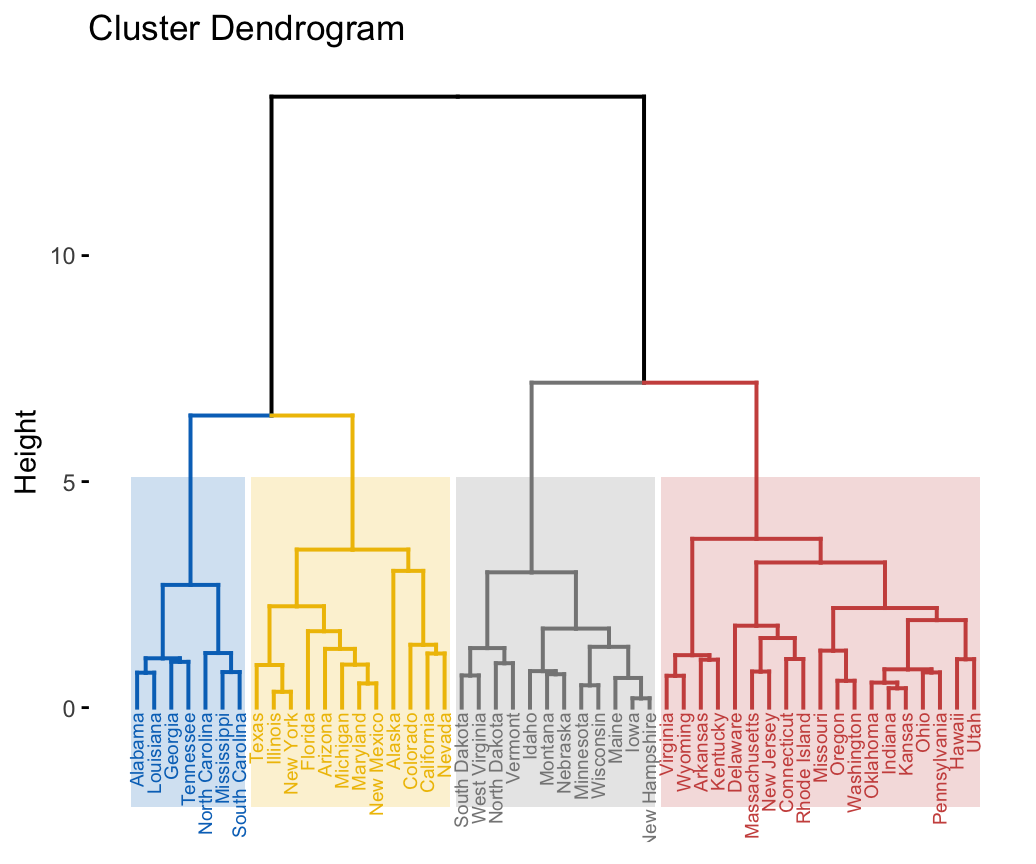
- Compare two dendrograms:
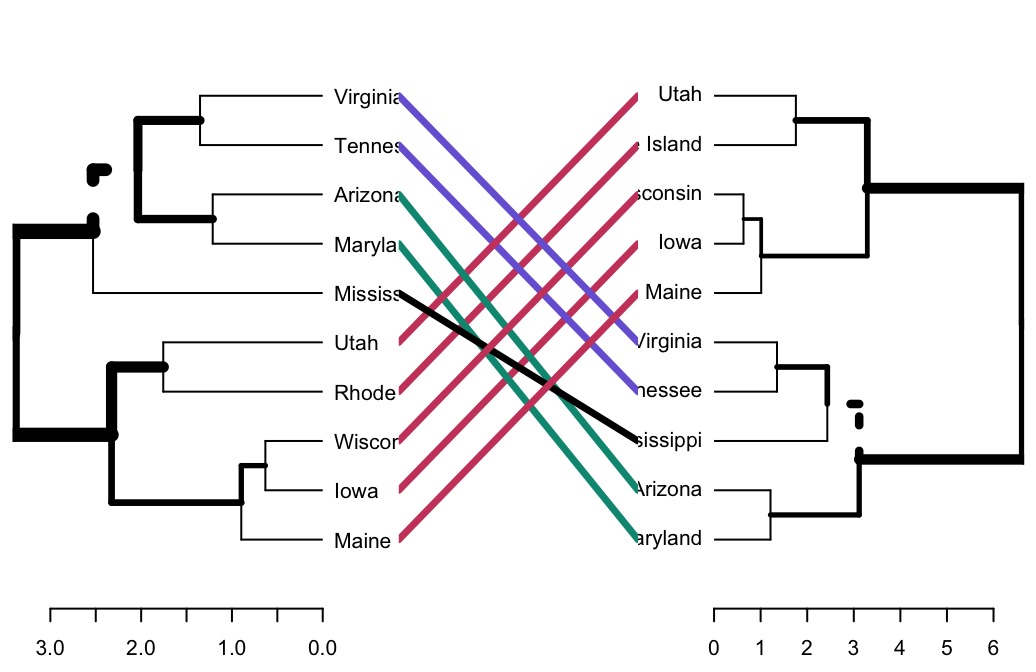
- Heatmap:
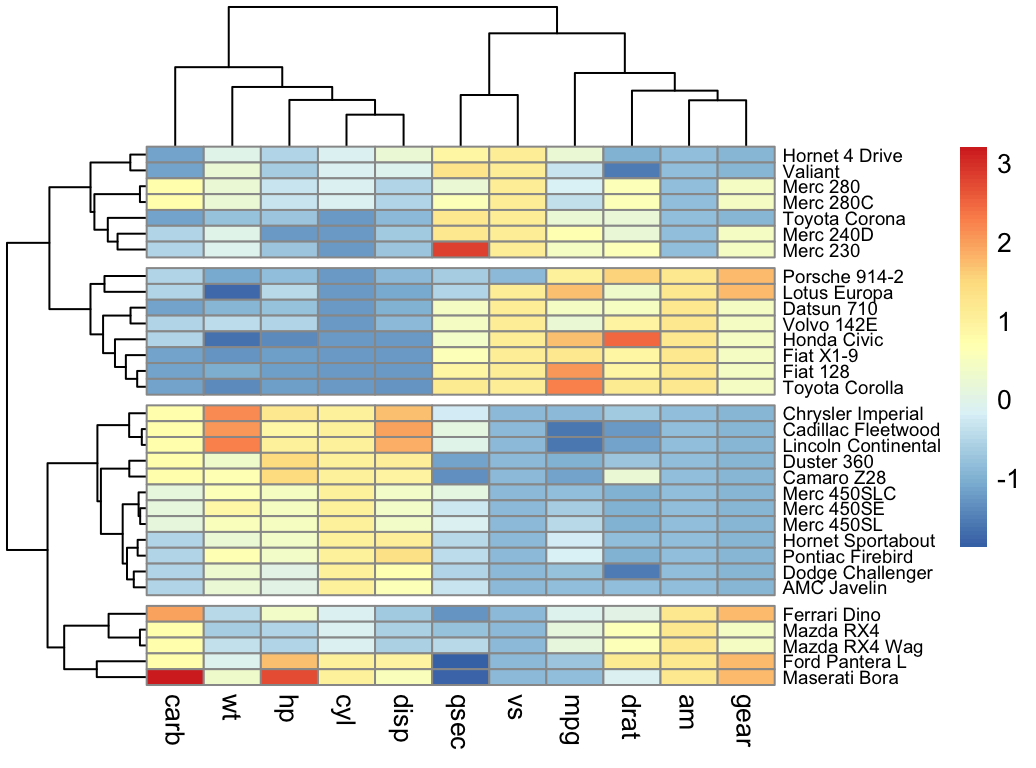
Part IV describes clustering validation and evaluation strategies, which consists of measuring the goodness of clustering results. Before applying any clustering algorithm to a data set, the first thing to do is to assess the clustering tendency. That is, whether applying clustering is suitable for the data. If yes, then how many clusters are there. Next, you can perform hierarchical clustering or partitioning clustering (with a pre-specified number of clusters). Finally, you can use a number of measures, described in this chapter, to evaluate the goodness of the clustering results.
The different chapters included in part IV are organized as follow:
- Assessing clustering tendency (Chapter 11)
- Determining the optimal number of clusters (Chapter 12)
- Cluster validation statistics (Chapter 13)
- Choosing the best clustering algorithms (Chapter 14)
- Computing p-value for hierarchical clustering (Chapter 15)
In this section, you’ll learn how to create and interpret the plots hereafter.
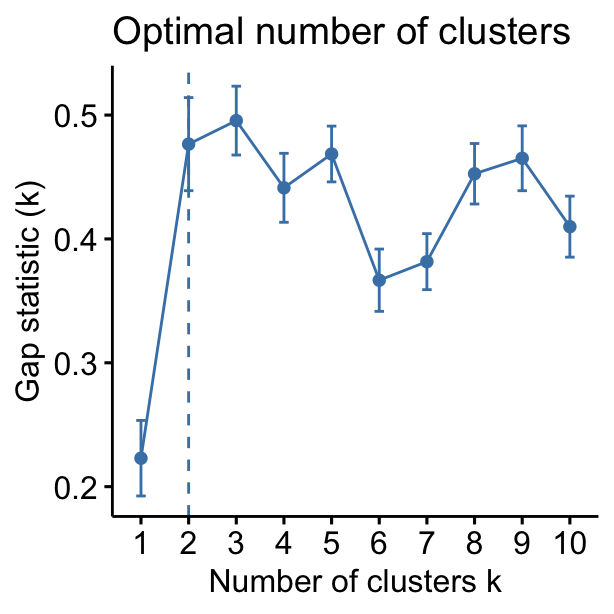
- Visual assessment of clustering tendency (left panel): Clustering tendency is detected in a visual form by counting the number of square shaped dark blocks along the diagonal in the image.
- Determine the optimal number of clusters (right panel) in a data set using the gap statistics.
## Clustering k = 1,2,..., K.max (= 10): .. done
## Bootstrapping, b = 1,2,..., B (= 100) [one "." per sample]:
## .................................................. 50
## .................................................. 100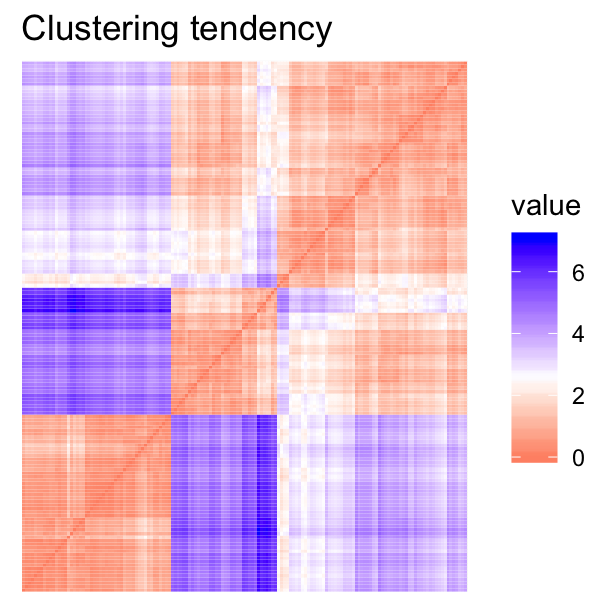
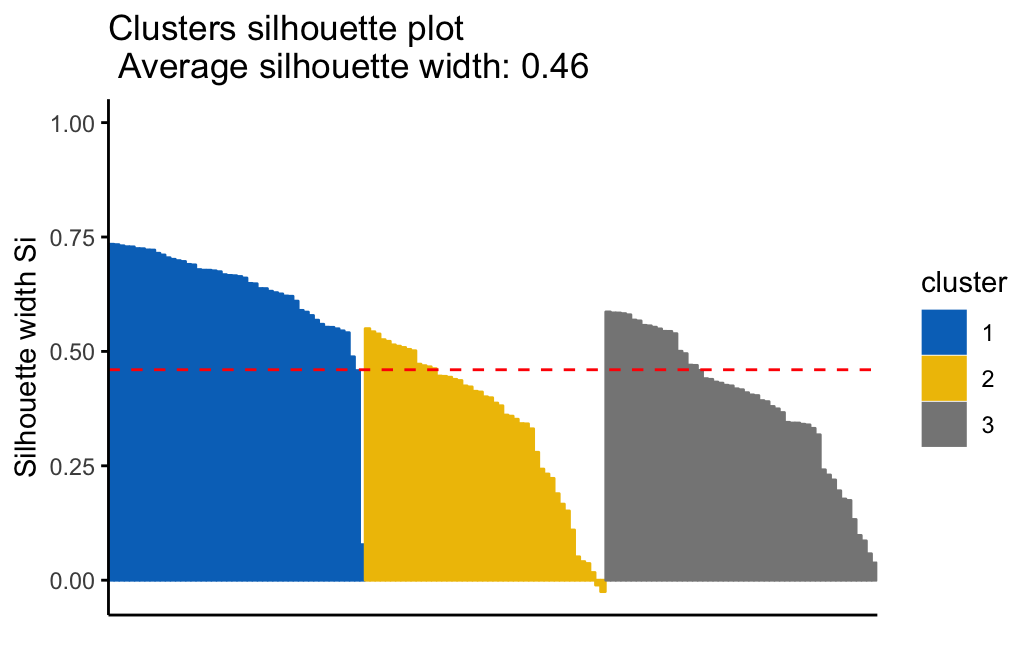
- Cluster validation using the silhouette coefficient (Si): A value of Si close to 1 indicates that the object is well clustered. A value of Si close to -1 indicates that the object is poorly clustered. The figure below shows the silhouette plot of a k-means clustering.
Part V presents advanced clustering methods, including:
- Hierarchical k-means clustering (Chapter 16)
- Fuzzy clustering (Chapter 17)
- Model-based clustering (Chapter 18)
- DBSCAN: Density-Based Clustering (Chapter 19)
The hierarchical k-means clustering is an hybrid approach for improving k-means results.
In Fuzzy clustering, items can be a member of more than one cluster. Each item has a set of membership coefficients corresponding to the degree of being in a given cluster.
In model-based clustering, the data are viewed as coming from a distribution that is mixture of two ore more clusters. It finds best fit of models to data and estimates the number of clusters.
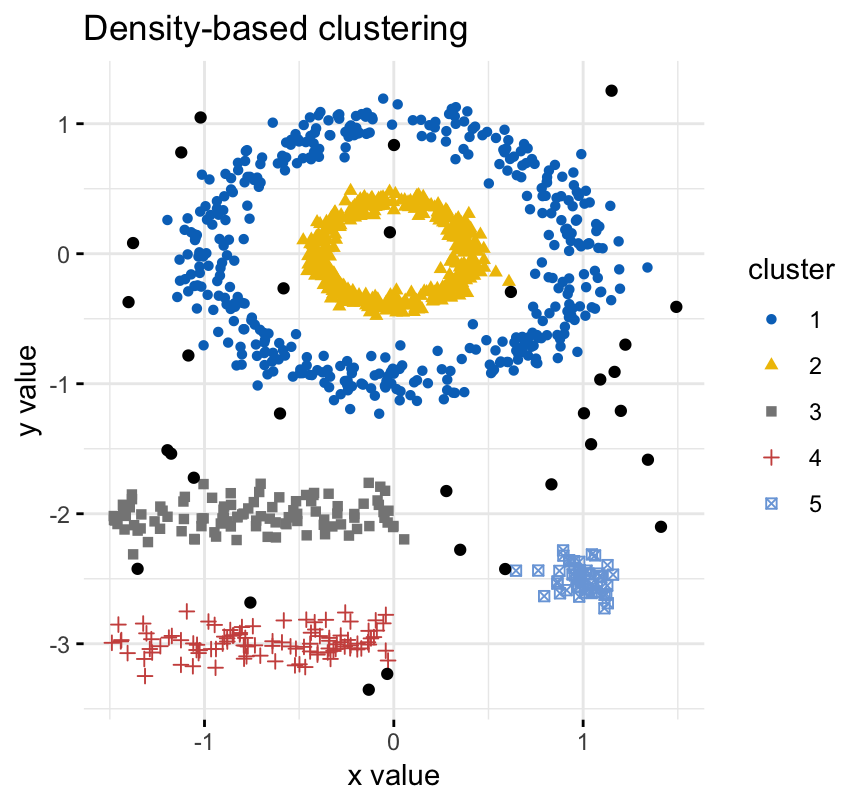
The density-based clustering (DBSCAN is a partitioning method that has been introduced in Ester et al. (1996). It can find out clusters of different shapes and sizes from data containing noise and outliers.
Recommended for you
This section contains best data science and self-development resources to help you on your path.
Coursera - Online Courses and Specialization
Data science
- Course: Machine Learning: Master the Fundamentals by Stanford
- Specialization: Data Science by Johns Hopkins University
- Specialization: Python for Everybody by University of Michigan
- Courses: Build Skills for a Top Job in any Industry by Coursera
- Specialization: Master Machine Learning Fundamentals by University of Washington
- Specialization: Statistics with R by Duke University
- Specialization: Software Development in R by Johns Hopkins University
- Specialization: Genomic Data Science by Johns Hopkins University
Popular Courses Launched in 2020
- Google IT Automation with Python by Google
- AI for Medicine by deeplearning.ai
- Epidemiology in Public Health Practice by Johns Hopkins University
- AWS Fundamentals by Amazon Web Services
Trending Courses
- The Science of Well-Being by Yale University
- Google IT Support Professional by Google
- Python for Everybody by University of Michigan
- IBM Data Science Professional Certificate by IBM
- Business Foundations by University of Pennsylvania
- Introduction to Psychology by Yale University
- Excel Skills for Business by Macquarie University
- Psychological First Aid by Johns Hopkins University
- Graphic Design by Cal Arts
Amazon FBA
Amazing Selling Machine
Books - Data Science
Our Books
- Practical Guide to Cluster Analysis in R by A. Kassambara (Datanovia)
- Practical Guide To Principal Component Methods in R by A. Kassambara (Datanovia)
- Machine Learning Essentials: Practical Guide in R by A. Kassambara (Datanovia)
- R Graphics Essentials for Great Data Visualization by A. Kassambara (Datanovia)
- GGPlot2 Essentials for Great Data Visualization in R by A. Kassambara (Datanovia)
- Network Analysis and Visualization in R by A. Kassambara (Datanovia)
- Practical Statistics in R for Comparing Groups: Numerical Variables by A. Kassambara (Datanovia)
- Inter-Rater Reliability Essentials: Practical Guide in R by A. Kassambara (Datanovia)
Others
- R for Data Science: Import, Tidy, Transform, Visualize, and Model Data by Hadley Wickham & Garrett Grolemund
- Hands-On Machine Learning with Scikit-Learn, Keras, and TensorFlow: Concepts, Tools, and Techniques to Build Intelligent Systems by Aurelien Géron
- Practical Statistics for Data Scientists: 50 Essential Concepts by Peter Bruce & Andrew Bruce
- Hands-On Programming with R: Write Your Own Functions And Simulations by Garrett Grolemund & Hadley Wickham
- An Introduction to Statistical Learning: with Applications in R by Gareth James et al.
- Deep Learning with R by François Chollet & J.J. Allaire
- Deep Learning with Python by François Chollet
Version:
 Français
Français






Eko Subagyo (verified owner) –
Hamidou Sy (verified owner) –
Bengt (verified owner) –
Excellent, easy to follow.
Christian Larsen (verified owner) –
Alex Sanchez Pla (verified owner) –
The book is both clear and complete and commands are straightforward to use.
It would be good to have a github repository to avid copy-paste
Anonymous (verified owner) –
Manuel Pellicer (verified owner) –
Johann (verified owner) –
A good book for people who have a basic understanding of cluster analysis and want to learn more, particularly the execution thereof within R. Clear graphics and good level of explanation. The book focusses on implementation and understanding of the methods, without having to struggle through pages of mathematical proofs, which some readers will appreciate.
Anonymous (verified owner) –
Anonymous (verified owner) –
Ognjen Žarić (verified owner) –
Chanvoleak (verified owner) –
Really help me to improve my knowledge on R
Modeste M. (verified owner) –
Very interesting book. It provides practical guide to cluster analysis, elegant visualization and interpretation. I recommend it.
Charles Abromaitis (verified owner) –
Bitrus (verified owner) –
very excellent book; highly recommended
RUBEN ADAD (verified owner) –
Andres Felipe Gavilan Orozco (verified owner) –
super helpfull book
Pavel (verified owner) –
Very professional level, very helpfull book
Anonymous (verified owner) –
Anonymous (verified owner) –
Andre S. (verified owner) –
Kaye (verified owner) –
So far, so good. Easy to understand and good illustrations.
Karol L. (verified owner) –
Perfect addition to your data science library
Jean-Carlos Montero-Serrano (verified owner) –
Excellent book ! but help is missing to make triangle diagrams, boxplot and bar plot with clusters (including cluster center).
Roberto Gil-Saura (verified owner) –
Thank you Mr. Kassambara for this simple, but useful guide. The way the analysis is approached is very accurate and easy to follow.
Anonymous (verified owner) –
Jhon Jairo Vargas Sánchez (verified owner) –
Excellent book. There are some digital errors and some number of equations are not visible
Etienne Ntumba (verified owner) –
Grzegorz Kulczycki (verified owner) –
Anonymous (verified owner) –
Found the answers to my questions in this book.
Saddam –
Can we get the R script file for each chapter?
SEBASTIÁN RIQUELME (verified owner) –
After purchasing the book, I got the link to download it as soon as I paid, so the service was good. The content is pivotal for my thesis project in my master studies.
michael frimpong (verified owner) –
Very good experience
Elisa Duarte (verified owner) –
Sílvia Panzo (verified owner) –
Anonymous (verified owner) –
Alexandre de Oliveira (verified owner) –
Clear explanations both of theory and codes
Nathalie M. (verified owner) –
Great book!
Cristian Opariuc-Dan (verified owner) –